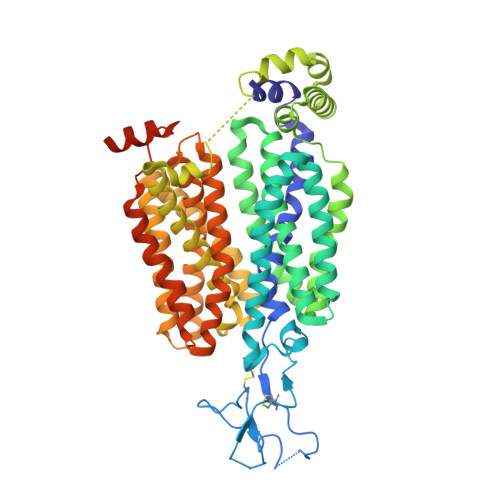
The substrate and inhibitor binding mechanism of polyspecific transporter OAT1 revealed by high-resolution cryo-EM.
Dou, T., Lian, T., Shu, S., He, Y., Jiang, J.(2023) Nat Struct Mol Biol 30: 1794-1805
- PubMed: 37845412 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01123-3
- Primary Citation Related Structures:
8SDU, 8SDY, 8SDZ - PubMed Abstract:
Organic anion transporters (OATs) of the SLC22 family have crucial roles in the transport of organic anions, including metabolites and therapeutic drugs, and in transporter-mediated drug-drug interactions. In the kidneys, OATs facilitate the elimination of metabolic waste products and xenobiotics. However, their transport activities can lead to the accumulation of certain toxic compounds within cells, causing kidney damage. Moreover, OATs are important drug targets, because their inhibition modulates the elimination or retention of substrates linked to diseases. Despite extensive research on OATs, the molecular basis of their substrate and inhibitor binding remains poorly understood. Here we report the cryo-EM structures of rat OAT1 (also known as SLC22A6) and its complexes with para-aminohippuric acid and probenecid at 2.1, 2.8 and 2.9 Å resolution, respectively. Our findings reveal a highly conserved substrate binding mechanism for SLC22 transporters, wherein four aromatic residues form a cage to accommodate the polyspecific binding of diverse compounds.
- Laboratory of Membrane Proteins and Structural Biology, Biochemistry and Biophysics Center, National Heart, Lung, and Blood Institute, Bethesda, MD, USA.
Organizational Affiliation: