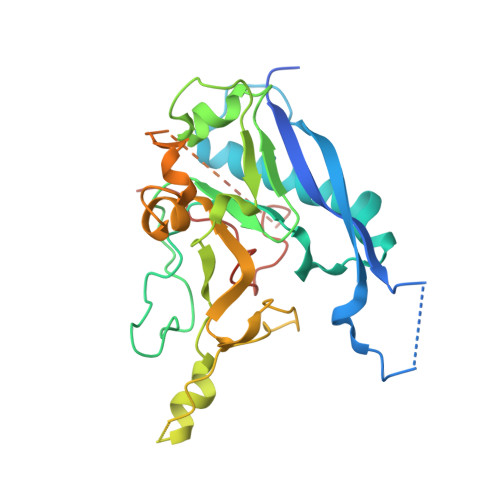
Engineering the ADDobody protein scaffold for generation of high-avidity ADDomer super-binders.
Buzas, D., Sun, H., Toelzer, C., Yadav, S.K.N., Borucu, U., Gautam, G., Gupta, K., Bufton, J.C., Capin, J., Sessions, R.B., Garzoni, F., Berger, I., Schaffitzel, C.(2024) Structure 32: 342
- PubMed: 38198950 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.12.010
- Primary Citation Related Structures:
8COI, 8QB3, 8QBX - PubMed Abstract:
Adenovirus-derived nanoparticles (ADDomer) comprise 60 copies of adenovirus penton base protein (PBP). ADDomer is thermostable, rendering the storage, transport, and deployment of ADDomer-based therapeutics independent of a cold chain. To expand the scope of ADDomers for new applications, we engineered ADDobodies, representing PBP crown domain, genetically separated from PBP multimerization domain. We inserted heterologous sequences into hyper-variable loops, resulting in monomeric, thermostable ADDobodies expressed at high yields in Escherichia coli. The X-ray structure of an ADDobody prototype validated our design. ADDobodies can be used in ribosome display experiments to select a specific binder against a target, with an enrichment factor of ∼10 4 -fold per round. ADDobodies can be re-converted into ADDomers by genetically reconnecting the selected ADDobody with the PBP multimerization domain from a different species, giving rise to a multivalent nanoparticle, called Chimera, confirmed by a 2.2 Å electron cryo-microscopy structure. Chimera comprises 60 binding sites, resulting in ultra-high, picomolar avidity to the target.
- School of Biochemistry, University of Bristol, University Walk, Bristol BS8 1TD, UK; Max Planck Bristol Centre for Minimal Biology, Cantock's Close, Bristol BS8 1TS, UK.
Organizational Affiliation: