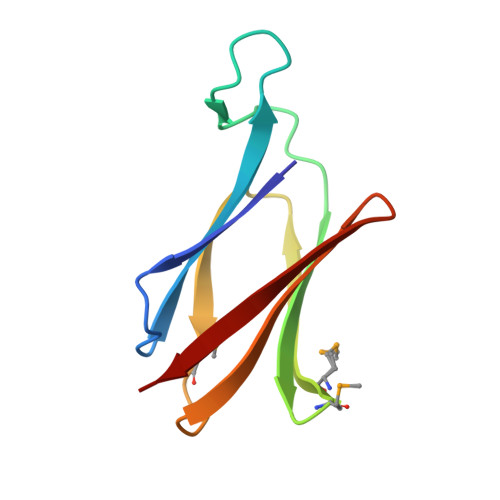
Structural and functional insights into the C-terminal signal domain of the Bacteroidetes type-IX secretion system.
Mizgalska, D., Rodriguez-Banqueri, A., Veillard, F., Ksiazek, M., Goulas, T., Guevara, T., Eckhard, U., Potempa, J., Gomis-Ruth, F.X.(2024) Open Biol 14: 230448-230448
- PubMed: 38862016 Search on PubMed
- DOI: https://doi.org/10.1098/rsob.230448
- Primary Citation Related Structures:
8QB1 - PubMed Abstract:
Gram-negative bacteria from the Bacteroidota phylum possess a type-IX secretion system (T9SS) for protein secretion, which requires cargoes to have a C-terminal domain (CTD). Structurally analysed CTDs are from Porphyromonas gingivalis proteins RgpB, HBP35, PorU and PorZ, which share a compact immunoglobulin-like antiparallel 3+4 β-sandwich (β1-β7). This architecture is essential as a P. gingivalis strain with a single-point mutant of RgpB disrupting the interaction of the CTD with its preceding domain prevented secretion of the protein. Next, we identified the C-terminus ('motif C-t.') and the loop connecting strands β3 and β4 ('motif Lβ3β4') as conserved. We generated two strains with insertion and replacement mutants of PorU, as well as three strains with ablation and point mutants of RgpB, which revealed both motifs to be relevant for T9SS function. Furthermore, we determined the crystal structure of the CTD of mirolase, a cargo of the Tannerella forsythia T9SS , which shares the same general topology as in Porphyromonas CTDs. However, motif Lβ3β4 was not conserved. Consistently, P. gingivalis could not properly secrete a chimaeric protein with the CTD of peptidylarginine deiminase replaced with this foreign CTD. Thus, the incompatibility of the CTDs between these species prevents potential interference between their T9SSs.
- Department of Microbiology, Faculty of Biochemistry, Biophysics and Biotechnology, Jagiellonian University, Kraków, Poland.
Organizational Affiliation: