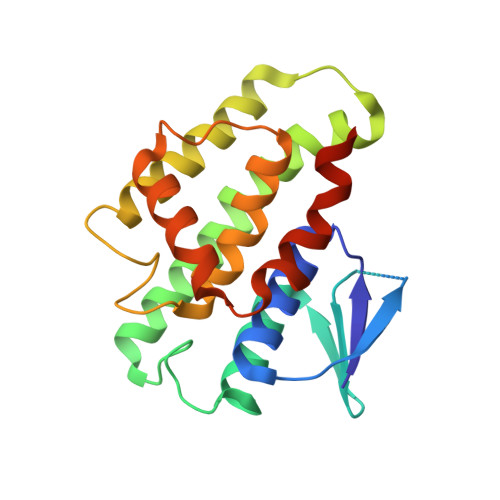
Structure-activity analysis suggests an olfactory function for the unique antennal delta glutathione transferase of Apis mellifera.
Schwartz, M., Boichot, V., Muradova, M., Fournier, P., Senet, P., Nicolai, A., Canon, F., Lirussi, F., Ladeira, R., Maibeche, M., Chertemps, T., Aubert, E., Didierjean, C., Neiers, F.(2023) FEBS Lett 597: 3038-3048
- PubMed: 37933500 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.14770
- Primary Citation Related Structures:
8Q89, 8Q8A, 8Q8B - PubMed Abstract:
Glutathione transferases (GST) are detoxification enzymes that conjugate glutathione to a wide array of molecules. In the honey bee Apis mellifera, AmGSTD1 is the sole member of the delta class of GSTs, with expression in antennae. Here, we structurally and biochemically characterized AmGSTD1 to elucidate its function. We showed that AmGSTD1 can efficiently catalyse the glutathione conjugation of classical GST substrates. Additionally, AmGSTD1 exhibits binding properties with a range of odorant compounds. AmGSTD1 has a peculiar interface with a structural motif we propose to call 'sulfur sandwich'. This motif consists of a cysteine disulfide bridge sandwiched between the sulfur atoms of two methionine residues and is stabilized by CH…S hydrogen bonds and S…S sigma-hole interactions. Thermal stability studies confirmed that this motif is important for AmGSTD1 stability and, thus, could facilitate its functions in olfaction.
- CSGA, Flavour Perception: Molecular Mechanisms (Flavours), Université de Bourgogne, INRAE, CNRS, Institut Agro, Dijon, France.
Organizational Affiliation: