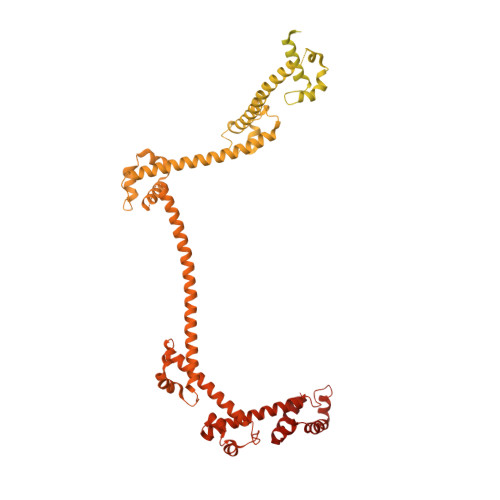
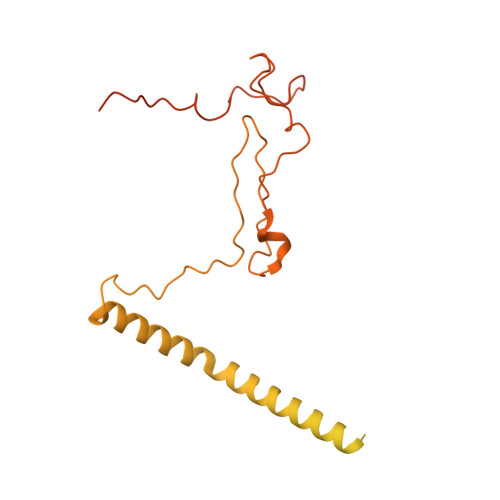
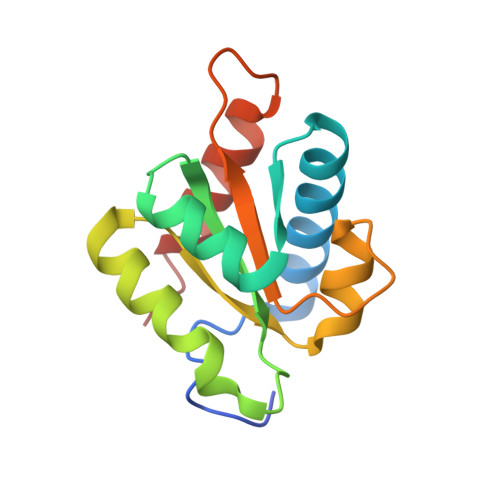
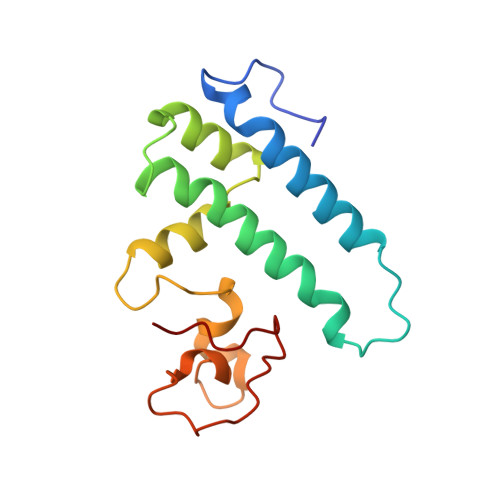
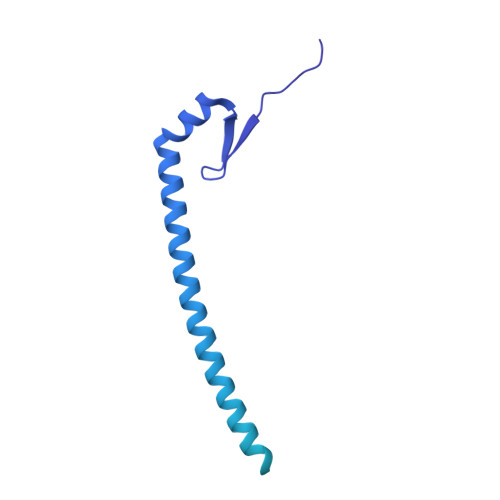
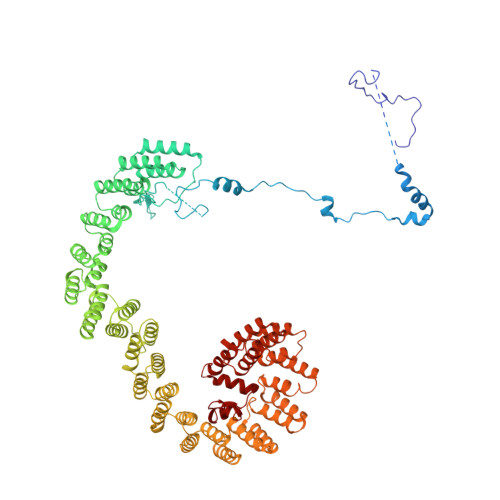
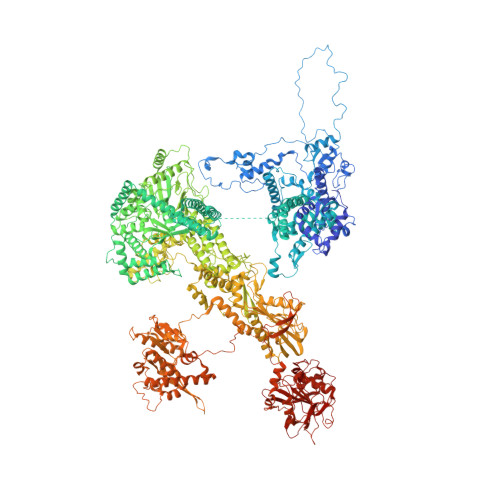
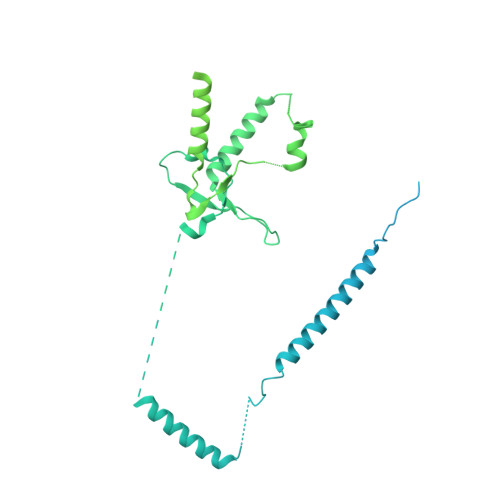
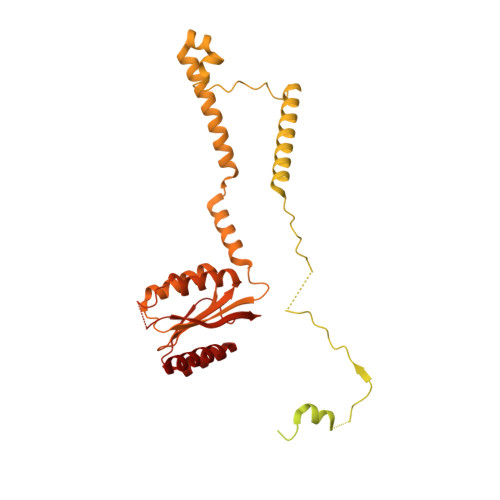
Cryo-EM analyses of dimerized spliceosomes provide new insights into the functions of B complex proteins.
Zhang, Z., Kumar, V., Dybkov, O., Will, C.L., Urlaub, H., Stark, H., Luhrmann, R.(2024) EMBO J 43: 1065-1088
- PubMed: 38383864 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-024-00052-1
- Primary Citation Related Structures:
8Q7N, 8QO9 - PubMed Abstract:
The B complex is a key intermediate stage of spliceosome assembly. To improve the structural resolution of monomeric, human spliceosomal B (hB) complexes and thereby generate a more comprehensive hB molecular model, we determined the cryo-EM structure of B complex dimers formed in the presence of ATP S. The enhanced resolution of these complexes allows a finer molecular dissection of how the 5' splice site (5'ss) is recognized in hB, and new insights into molecular interactions of FBP21, SNU23 and PRP38 with the U6/5'ss helix and with each other. It also reveals that SMU1 and RED are present as a heterotetrameric complex and are located at the interface of the B dimer protomers. We further show that MFAP1 and UBL5 form a 5' exon binding channel in hB, and elucidate the molecular contacts stabilizing the 5' exon at this stage. Our studies thus yield more accurate models of protein and RNA components of hB complexes. They further allow the localization of additional proteins and protein domains (such as SF3B6, BUD31 and TCERG1) whose position was not previously known, thereby uncovering new functions for B-specific and other hB proteins during pre-mRNA splicing.
- Department of Structural Dynamics, Max-Planck-Institute for Multidisciplinary Sciences, Am Fassberg 11, 37077, Göttingen, Germany.
Organizational Affiliation: