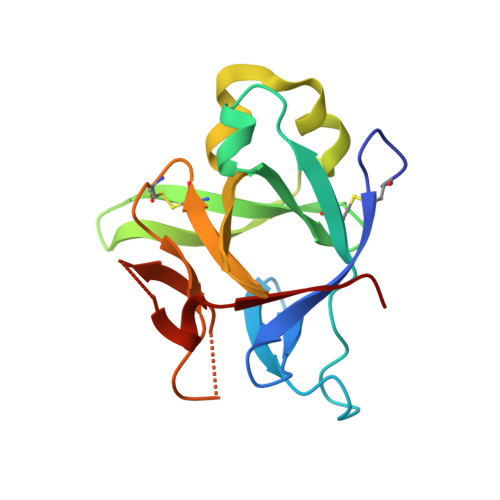
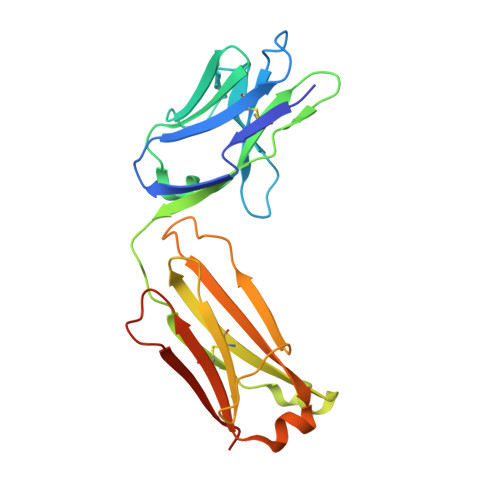
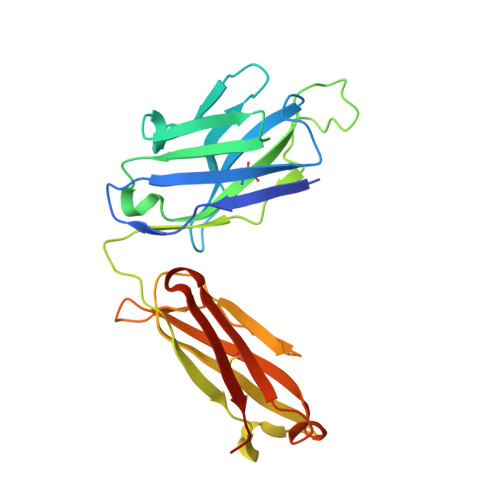
Neutralizing IL-38 activates gamma delta T cell-dependent antitumor immunity and sensitizes for chemotherapy.
da Silva, P., Mora, J., You, X., Wiechmann, S., Putyrski, M., Garcia-Pardo, J., Kannt, A., Ernst, A., Bruene, B., Weigert, A.(2024) J Immunother Cancer 12
- PubMed: 39209451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1136/jitc-2023-008641
- Primary Citation Related Structures:
8Q3J - PubMed Abstract:
The interleukin (IL)-1-family receptor antagonist IL-38 has emerged as a negative regulator of auto-inflammation. Given the intricate interplay between antitumor immunity and auto-inflammation, we hypothesized that blocking IL-38 may enhance tumor immune control. Our hypothesis was tested in the transgenic polyoma virus middle T oncoprotein mammary carcinoma model that is suitable for identifying strong immunomodulators. To investigate the effect of acute IL-38 blockade, we used a neutralizing antibody, alone or in combination with chemotherapy. Immune cell composition and location in tumors were determined by flow cytometry and immunohistochemistry, respectively. The role of γδ T cells was studied using an antibody blocking γδ T-cell receptor signaling. Whole transcriptome RNA sequencing and RNA expression analysis were employed to determine mechanisms downstream of IL-38 neutralization. Additionally, in vitro assays with γδ T cells, CD8+ T cells and cDC1, followed by in vivo CD8+ T cell depletion, were performed to study the underlying mechanistic pathways. Both, genetic ablation of IL-38 and neutralization with the antibody, reduced tumorigenesis, and IL-38 blockade improved chemotherapy efficacy. This was accompanied by an augmented lymphocyte infiltrate dominated by γδ T cells and CD8+ T cells, and signaling through the γδ-T-cell receptor was required for CD8+ T cell infiltration. Rather than directly interacting with CD8+ T cells, γδ T cells recruited conventional dendritic cells (cDC1) into tumors via the chemokine Xcl1. cDC1 in turn activated CD8+ T cells via the Notch pathway. Moreover, IL-38 negatively correlated with cDC1, XCL1-producing γδ T cells, T-cell infiltrates and survival in patients with mammary carcinoma. These data suggest that interfering with IL-38 improves antitumor immunity even in immunologically cold tumors.
- Faculty of Medicine, Institute of Biochemistry I, Goethe-University Frankfurt, Frankfurt, Germany.
Organizational Affiliation: