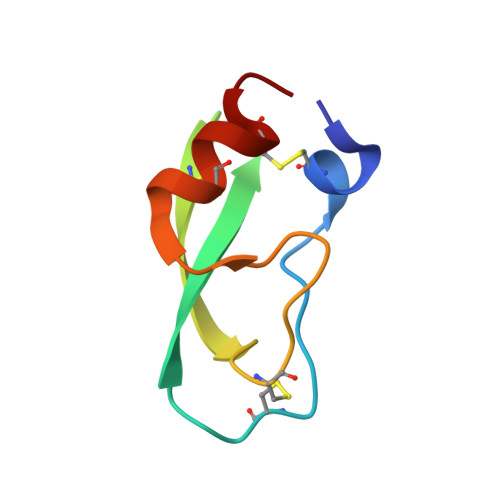
Crystal structure of a Y35G mutant of bovine pancreatic trypsin inhibitor.
Housset, D., Kim, K.S., Fuchs, J., Woodward, C., Wlodawer, A.(1991) J Mol Biology 220: 757-770
- PubMed: 1714504 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90115-m
- Primary Citation Related Structures:
8PTI - PubMed Abstract:
The structure of a Y35G mutant of bovine pancreatic trypsin inhibitor (BPTI) was solved by molecular replacement and was refined by both simulated annealing and restrained least-squares at 1.8 A resolution. The crystals belong to the space group P42212, with unit cell dimensions a = b = 46.75 A, c = 50.61 A. The final R-factor is 0.159 and the deviation from ideality for bond distances is 0.02 A. The structure of the mutant differs from that of the native protein, showing an overall root-mean-square (r.m.s.) difference of 1.86 A for main-chain atoms. However, the change is mostly localized in the two loops (respective r.m.s. values of 2.04 A and 3.93 A) and the C terminus (r.m.s. 6.79 A), while the core of the protein is well conserved (r.m.s. 0.45 A). The change in the loop regions can be clearly attributed to the mutation while the difference in the C terminus might be only due to a different crystal packing. Seventy water molecules were included in the model but only seven of them are shared with the native structure. Thermal parameters are showing a good correlation with those for the wild-type of BPTI.
- Macromolecular Structure Laboratory, NCI-Frederick Cancer Research and Development Center, MD 21702.
Organizational Affiliation: