Computational Prediction of Cyclic Peptide Structural Ensembles and Application to the Design of Keap1 Binders.
Fonseca Lopez, F., Miao, J., Damjanovic, J., Bischof, L., Braun, M.B., Ling, Y., Hartmann, M.D., Lin, Y.S., Kritzer, J.A.(2023) J Chem Inf Model 63: 6925-6937
- PubMed: 37917529 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jcim.3c01337
- Primary Citation Related Structures:
8PKU, 8PKV, 8PKW, 8PKX - PubMed Abstract:
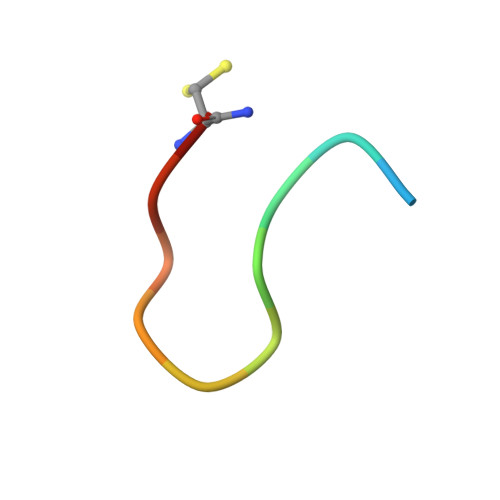
The Nrf2 transcription factor is a master regulator of the cellular response to oxidative stress, and Keap1 is its primary negative regulator. Activating Nrf2 by inhibiting the Nrf2-Keap1 protein-protein interaction has shown promise for treating cancer and inflammatory diseases. A loop derived from Nrf2 has been shown to inhibit Keap1 selectively, especially when cyclized, but there are no reliable design methods for predicting an optimal macrocyclization strategy. In this work, we employed all-atom, explicit-solvent molecular dynamics simulations with enhanced sampling methods to predict the relative degree of preorganization for a series of peptides cyclized with a set of bis-thioether "staples". We then correlated these predictions to experimentally measured binding affinities for Keap1 and crystal structures of the cyclic peptides bound to Keap1. This work showcases a computational method for designing cyclic peptides by simulating and comparing their entire solution-phase ensembles, providing key insights into designing cyclic peptides as selective inhibitors of protein-protein interactions.
- Department of Chemistry, Tufts University, Medford, Massachusetts 02155, United States.
Organizational Affiliation: