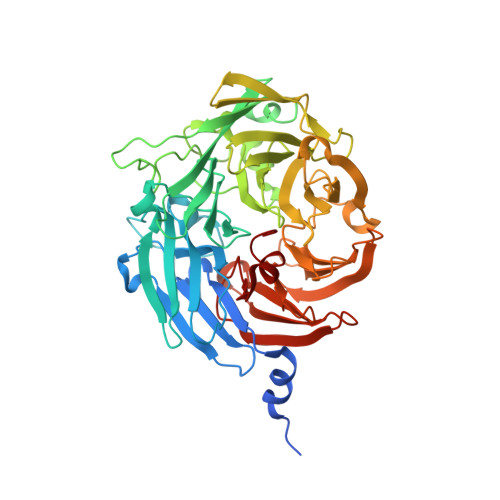
Improvement of the Diffraction Properties of Thiocyanate Dehydrogenase Crystals
Varfolomeeva, L.A., Polyakov, K.M., Komolov, A.S., Rakitina, T.V., Dergousova, N.I., Dorovatovskii, P.V., Boyko, K.M., Tikhonova, T.V., Popov, V.O.(2023) Crystallogr Rep