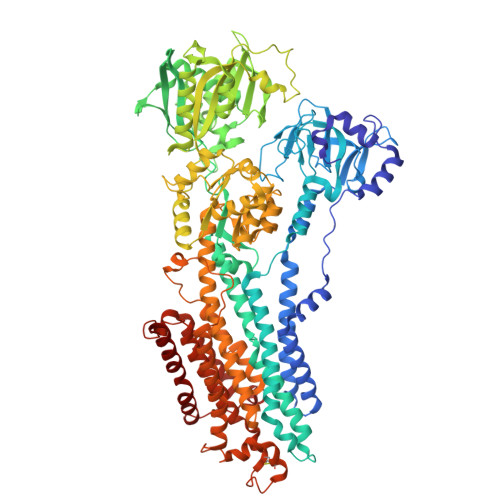
Photoswitchable inhibitors of the sarco(endo)plasmic calcium pump
Hjorth-Jensen, S.J., Konrad, D.B., Quistgaard, E.M.H., Hansen, L.C., Novak, A., Chu, H., Jurasek, M., Zimmermann, T., Andersen, J.L., Baran, P.S., Nissen, P., Trauner, D.To be published.