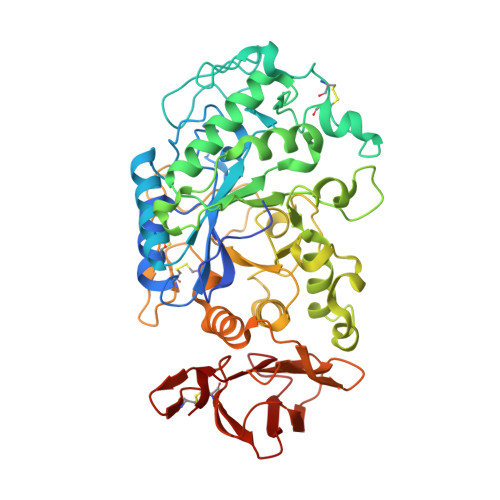
Structural and Functional Characterization of Drosophila melanogaster alpha-Amylase.
Rhimi, M., Da Lage, J.L., Haser, R., Feller, G., Aghajari, N.(2023) Molecules 28
- PubMed: 37513201 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules28145327
- Primary Citation Related Structures:
8OR6, 8ORP - PubMed Abstract:
Insects rely on carbohydrates such as starch and glycogen as an energy supply for growth of larvae and for longevity. In this sense α-amylases have essential roles under extreme conditions, e.g., during nutritional or temperature stress, thereby contributing to survival of the insect. This makes them interesting targets for combating insect pests. Drosophila melanogaster α-amylase, DMA, which belongs to the glycoside hydrolase family 13, sub family 15, has been studied from an evolutionary, biochemical, and structural point of view. Our studies revealed that the DMA enzyme is active over a broad temperature and pH range, which is in agreement with the fluctuating environmental changes with which the insect is confronted. Crystal structures disclosed a new nearly fully solvated metal ion, only coordinated to the protein via Gln263. This residue is only conserved in the subgroup of D. melanogaster and may thus contribute to the enzyme adaptive response to large temperature variations. Studies of the effect of plant inhibitors and the pseudo-tetrasaccharide inhibitor acarbose on DMA activity, allowed us to underline the important role of the so-called flexible loop on activity/inhibition, but also to suggest that the inhibition modes of the wheat inhibitors WI-1 and WI-3 on DMA, are likely different.
- Molecular Microbiology and Structural Biochemistry, UMR5086, CNRS, University of Lyon 1, 7 Passage du Vercors, F-69367 Lyon, CEDEX 07, France.
Organizational Affiliation: