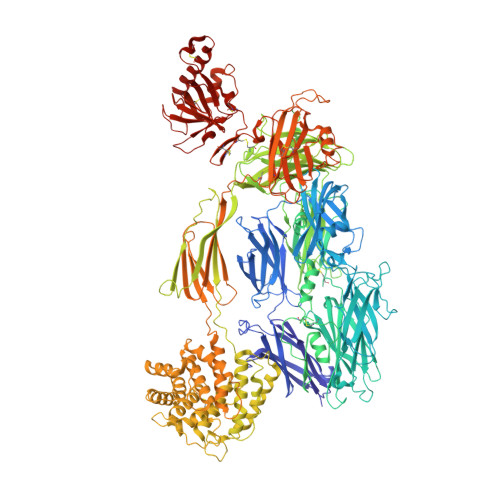
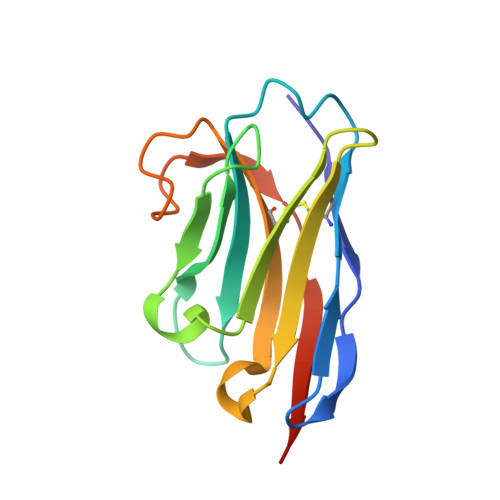
Cryo-EM analysis of complement C3 reveals a reversible major opening of the macroglobulin ring.
Gadeberg, T.A.F., Jorgensen, M.H., Olesen, H.G., Lorentzen, J., Harwood, S.L., Almeida, A.V., Fruergaard, M.U., Jensen, R.K., Kanis, P., Pedersen, H., Tranchant, E., Petersen, S.V., Thogersen, I.B., Kragelund, B.B., Lyons, J.A., Enghild, J.J., Andersen, G.R.(2025) Nat Struct Mol Biol
- PubMed: 39849196 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01467-4
- Primary Citation Related Structures:
8OQ3 - PubMed Abstract:
The C3 protein is the central molecule within the complement system and undergoes proteolytic activation to C3b in the presence of pathogens. Pattern-independent activation of C3 also occurs via hydrolysis, resulting in C3(H 2 O), but the structural details of C3 hydrolysis remain elusive. Here we show that the conformation of the C3(H 2 O) analog, C3MA, is indistinguishable from C3b. In contrast, the reaction intermediate C3* adopts a conformation dramatically different from both C3 and C3MA. In C3*, unlocking of the macroglobulin (MG) 3 domain creates a large opening in the MG ring through which the anaphylatoxin (ANA) domain translocates through a transient opening. C3MA formation is inhibited by an MG3-specific nanobody and prevented by linking the ANA domain to the C3 β-chain. Our study reveals an unexpected dynamic behavior of C3 and forms the basis for elucidation of the in vivo contribution of C3 hydrolysis and for controlling complement upon intravascular hemolysis and surface-contact-induced activation.
- Department of Molecular Biology and Genetics, Aarhus, Denmark.
Organizational Affiliation: