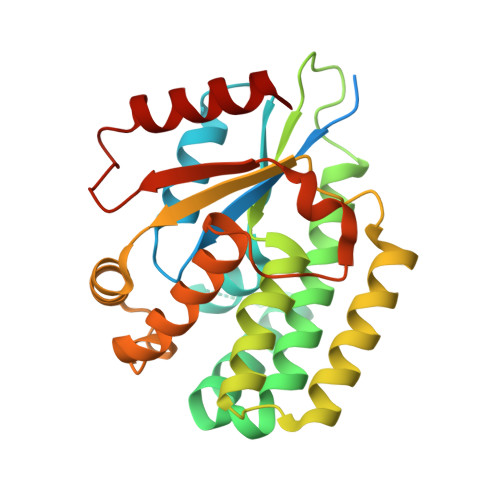
4'-Ethynyl-2'-Deoxycytidine (EdC) Preferentially Targets Lymphoma and Leukemia Subtypes by Inducing Replicative Stress.
Calbert, M.L., Chandramouly, G., Adams, C.M., Saez-Ayala, M., Kent, T., Tyagi, M., Ayyadevara, V.S.S.A., Wang, Y., Krais, J.J., Gordon, J., Atkins, J., Toma, M.M., Betzi, S., Boghossian, A.S., Rees, M.G., Ronan, M.M., Roth, J.A., Goldman, A.R., Gorman, N., Mitra, R., Childers, W.E., Grana, X., Skorski, T., Johnson, N., Hurtz, C., Morelli, X., Eischen, C.M., Pomerantz, R.T.(2024) Mol Cancer Ther 23: 683-699
- PubMed: 38064712 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-23-0487
- Primary Citation Related Structures:
8OOJ - PubMed Abstract:
Anticancer nucleosides are effective against solid tumors and hematologic malignancies, but typically are prone to nucleoside metabolism resistance mechanisms. Using a nucleoside-specific multiplexed high-throughput screening approach, we discovered 4'-ethynyl-2'-deoxycytidine (EdC) as a third-generation anticancer nucleoside prodrug with preferential activity against diffuse large B-cell lymphoma (DLBCL) and acute lymphoblastic leukemia (ALL). EdC requires deoxycytidine kinase (DCK) phosphorylation for its activity and induces replication fork arrest and accumulation of cells in S-phase, indicating it acts as a chain terminator. A 2.1Å cocrystal structure of DCK bound to EdC and UDP reveals how the rigid 4'-alkyne of EdC fits within the active site of DCK. Remarkably, EdC was resistant to cytidine deamination and SAMHD1 metabolism mechanisms and exhibited higher potency against ALL compared with FDA-approved nelarabine. Finally, EdC was highly effective against DLBCL tumors and B-ALL in vivo. These data characterize EdC as a preclinical nucleoside prodrug candidate for DLBCL and ALL.
- Department of Biochemistry and Molecular Biology, Sidney Kimmel Cancer Center, Thomas Jefferson University, Philadelphia, Pennsylvania.
Organizational Affiliation: