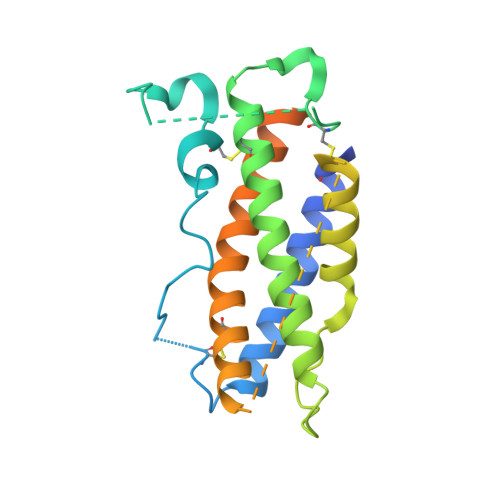
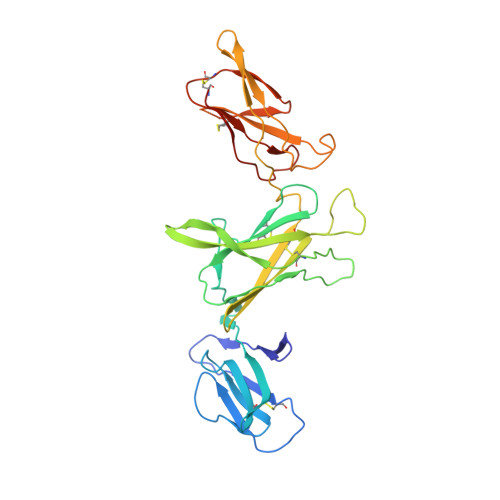
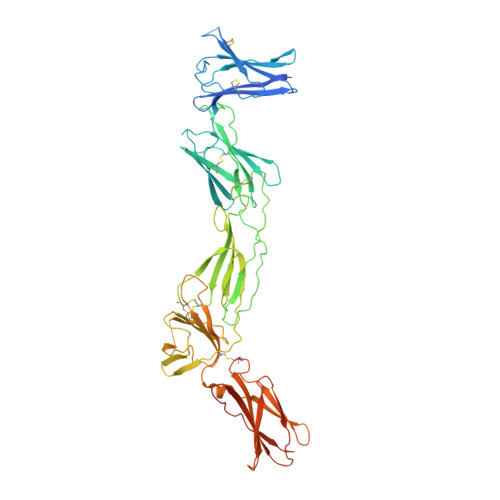
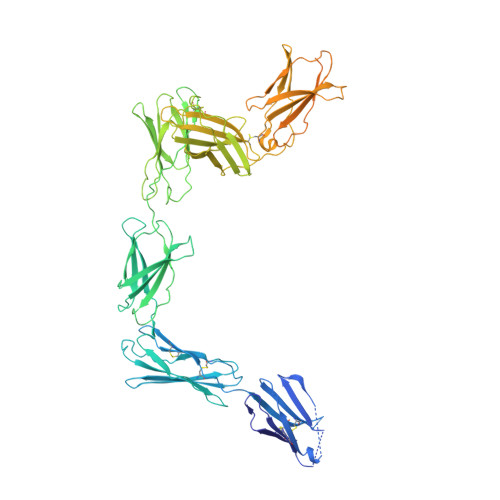
Structures of complete extracellular receptor assemblies mediated by IL-12 and IL-23.
Bloch, Y., Felix, J., Merceron, R., Provost, M., Symakani, R.A., De Backer, R., Lambert, E., Mehdipour, A.R., Savvides, S.N.(2024) Nat Struct Mol Biol 31: 591-597
- PubMed: 38287195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01190-6
- Primary Citation Related Structures:
8C7M, 8CR5, 8CR6, 8CR8, 8ODX, 8ODZ, 8OE0, 8OE4, 8PB1, 8PPM - PubMed Abstract:
Cell-surface receptor complexes mediated by pro-inflammatory interleukin (IL)-12 and IL-23, both validated therapeutic targets, are incompletely understood due to the lack of structural insights into their complete extracellular assemblies. Furthermore, there is a paucity of structural details describing the IL-12-receptor interaction interfaces, in contrast to IL-23-receptor complexes. Here we report structures of fully assembled mouse IL-12/human IL-23-receptor complexes comprising the complete extracellular segments of the cognate receptors determined by electron cryo-microscopy. The structures reveal key commonalities but also surprisingly diverse features. Most notably, whereas IL-12 and IL-23 both utilize a conspicuously presented aromatic residue on their α-subunit as a hotspot to interact with the N-terminal Ig domain of their high-affinity receptors, only IL-12 juxtaposes receptor domains proximal to the cell membrane. Collectively, our findings will help to complete our understanding of cytokine-mediated assemblies of tall cytokine receptors and will enable a cytokine-specific interrogation of IL-12/IL-23 signaling in physiology and disease.
- Unit for Structural Biology, Department of Biochemistry and Microbiology, Ghent University, Ghent, Belgium.
Organizational Affiliation: