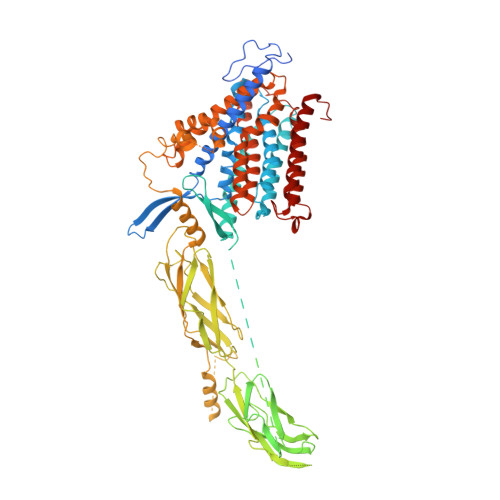
Structural insight into the allosteric inhibition of human sodium-calcium exchanger NCX1 by XIP and SEA0400.
Dong, Y., Yu, Z., Li, Y., Huang, B., Bai, Q., Gao, Y., Chen, Q., Li, N., He, L., Zhao, Y.(2024) EMBO J 43: 14-31
- PubMed: 38177313 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-023-00013-0
- Primary Citation Related Structures:
8JP0 - PubMed Abstract:
Sodium-calcium exchanger proteins influence calcium homeostasis in many cell types and participate in a wide range of physiological and pathological processes. Here, we elucidate the cryo-EM structure of the human Na + /Ca 2+ exchanger NCX1.3 in the presence of a specific inhibitor, SEA0400. Conserved ion-coordinating residues are exposed on the cytoplasmic face of NCX1.3, indicating that the observed structure is stabilized in an inward-facing conformation. We show how regulatory calcium-binding domains (CBDs) assemble with the ion-translocation transmembrane domain (TMD). The exchanger-inhibitory peptide (XIP) is trapped within a groove between the TMD and CBD2 and predicted to clash with gating helices TMs 1/6 at the outward-facing state, thus hindering conformational transition and promoting inactivation of the transporter. A bound SEA0400 molecule stiffens helix TM2ab and affects conformational rearrangements of TM2ab that are associated with the ion-exchange reaction, thus allosterically attenuating Ca 2+ -uptake activity of NCX1.3.
- Key Laboratory of Biomacromolecules (CAS), National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, 100101, China.
Organizational Affiliation: