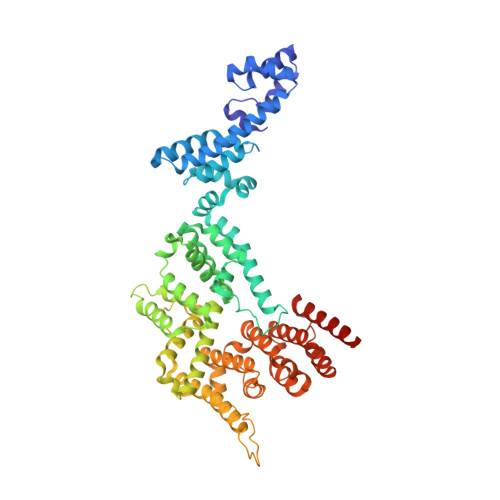
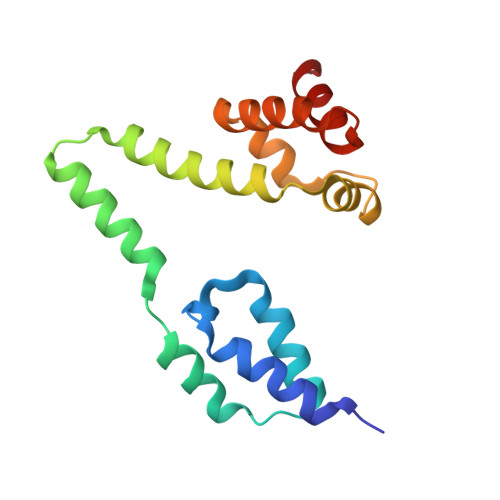
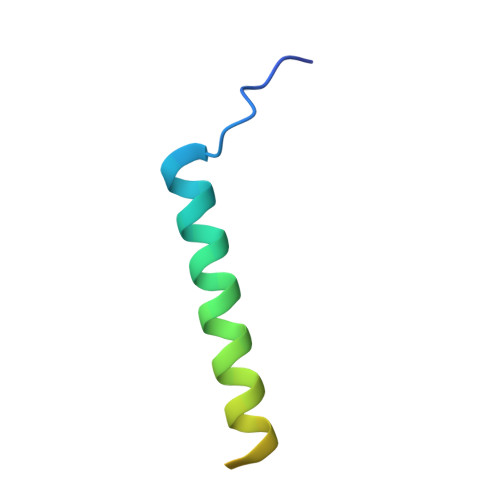
Structural basis of the human negative elongation factor NELF-B/C/E ternary complex.
Cao, Y., Qin, Y., Zhang, W., Tian, W., Ren, Y., Ren, J., Wang, J., Wang, M., Jiang, J., Wang, Z.(2023) Biochem Biophys Res Commun 677: 155-161
- PubMed: 37591184 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.08.019
- Primary Citation Related Structures:
8JJ6 - PubMed Abstract:
Negative elongation factor (NELF) is a four-subunit transcription elongation factor that mainly functions in maintaining the paused state of RNA polymerase II in eukaryotes. Upon binding to Pol II, NELF works synergistically with DRB sensitivity-inducing factor (DSIF) and inhibits transcription elongation of Pol II, which subsequently retains a stably paused state 20-60 base pairs downstream of the promoter. The promoter-proximal pausing of Pol II caused by NELF is a general mechanism of transcriptional regulation for most signal-responsive genes. To date, structural studies have significantly advanced our understanding of the molecular mechanisms of NELF. However, a high quality structural model clarifying the interaction details of this complex is still lacking. In this study, we solved the high resolution crystal structure of the NELF-B/C/E ternary complex. We observed detailed interactions between subunits and identified residues important for the association between NELF-B and NELF-E. Our work presents a precise model of the NELF complex, which will facilitate our understanding of its in vivo function.
- Key Laboratory of Cell Proliferation and Regulation Biology of Ministry of Education, College of Life Sciences, Beijing Normal University, 19 Xinjiekouwai Avenue, Beijing, 100875, China.
Organizational Affiliation: