A framework for Frizzled-G protein coupling and implications to the PCP signaling pathways.
Zhang, Z., Lin, X., Wei, L., Wu, Y., Xu, L., Wu, L., Wei, X., Zhao, S., Zhu, X., Xu, F.(2024) Cell Discov 10: 3-3
- PubMed: 38182578 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-023-00627-y
- Primary Citation Related Structures:
8J9N, 8J9O, 8JH7, 8JHB, 8JHC, 8JHI - PubMed Abstract:
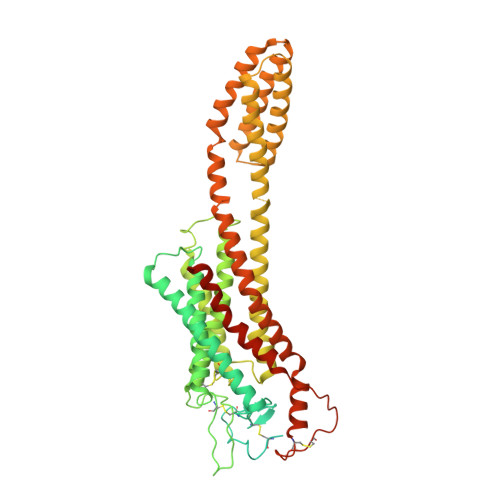
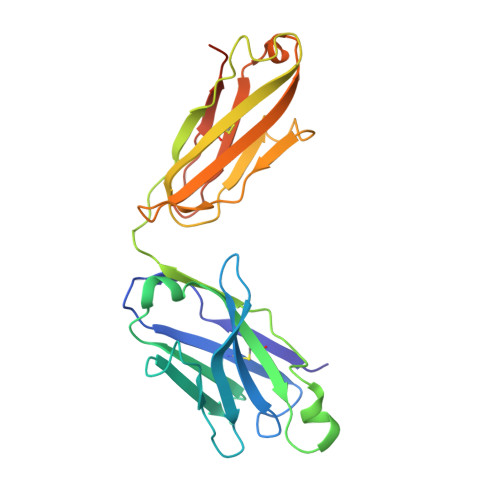
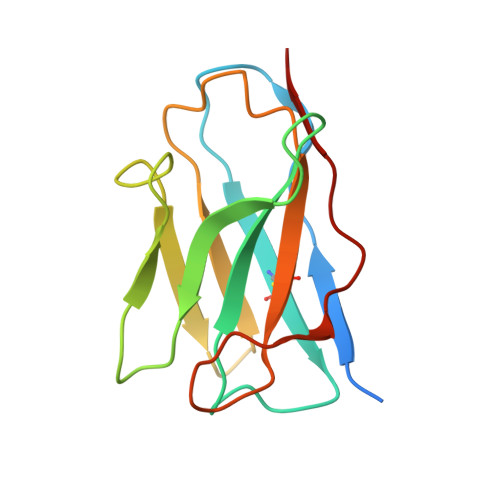
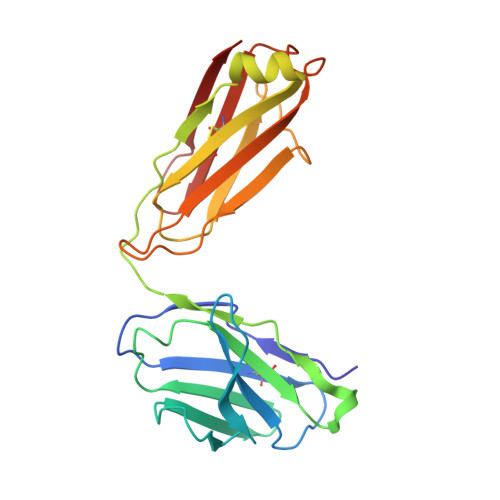
The ten Frizzled receptors (FZDs) are essential in Wnt signaling and play important roles in embryonic development and tumorigenesis. Among these, FZD6 is closely associated with lens development. Understanding FZD activation mechanism is key to unlock these emerging targets. Here we present the cryo-EM structures of FZD6 and FZD3 which are known to relay non-canonical planar cell polarity (PCP) signaling pathways as well as FZD1 in their G protein-coupled states and in the apo inactive states, respectively. Comparison of the three inactive/active pairs unveiled a shared activation framework among all ten FZDs. Mutagenesis along with imaging and functional analysis on the human lens epithelial tissues suggested potential crosstalk between the G-protein coupling of FZD6 and the PCP signaling pathways. Together, this study provides an integrated understanding of FZD structure and function, and lays the foundation for developing therapeutic modulators to activate or inhibit FZD signaling for a range of disorders including cancers and cataracts.
- iHuman Institute, ShanghaiTech University, Shanghai, China.
Organizational Affiliation: