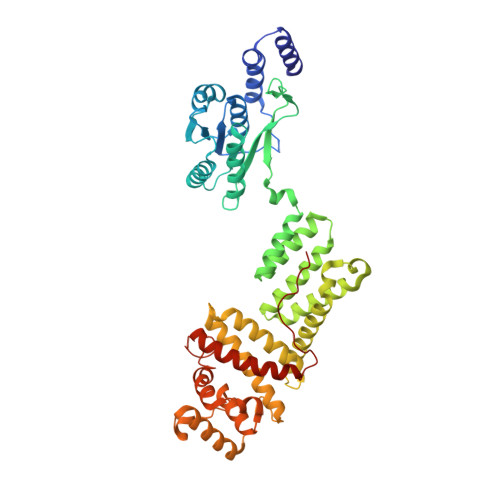
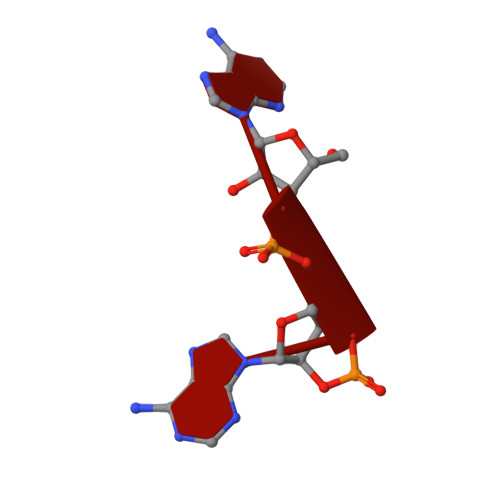
Molecular mechanism of allosteric activation of the CRISPR ribonuclease Csm6 by cyclic tetra-adenylate.
Du, L., Zhu, Q., Lin, Z.(2024) EMBO J 43: 304-315
- PubMed: 38177499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-023-00017-w
- Primary Citation Related Structures:
8JBB, 8JBC, 8JH1 - PubMed Abstract:
Type III CRISPR systems are innate immune systems found in bacteria and archaea, which produce cyclic oligoadenylate (cOA) second messengers in response to viral infections. In these systems, Csm6 proteins serve as ancillary nucleases that degrade single-stranded RNA (ssRNA) upon activation by cOA. In addition, Csm6 proteins also possess cOA-degrading activity as an intrinsic off-switch to avoid degradation of host RNA and DNA that would eventually lead to cell dormancy or cell death. Here, we present the crystal structures of Thermus thermophilus (Tt) Csm6 alone, and in complex with cyclic tetra-adenylate (cA 4 ) in both pre- and post-cleavage states. These structures establish the molecular basis of the long-range allosteric activation of TtCsm6 ribonuclease by cA 4 . cA 4 binding induces significant conformational changes, including closure of the CARF domain, dimerization of the HTH domain, and reorganization of the R-X 4-6 -H motif within the HEPN domain. The cleavage of cA 4 by the CARF domain restores each domain to a conformation similar to its apo state. Furthermore, we have identified hyperactive TtCsm6 variants that exhibit sustained cA 4 -activated RNase activity, showing great promise for their applications in genome editing and diagnostics.
- College of Chemistry, Fuzhou University, 350108, Fuzhou, China.
Organizational Affiliation: