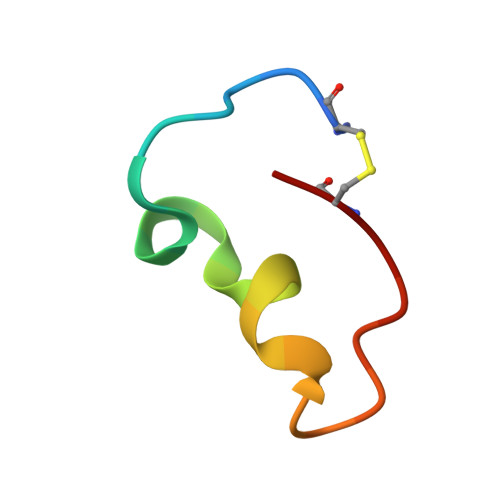
Machine learning and genetic algorithm-guided directed evolution for the development of antimicrobial peptides.
Zhang, H., Wang, Y., Zhu, Y., Huang, P., Gao, Q., Li, X., Chen, Z., Liu, Y., Jiang, J., Gao, Y., Huang, J., Qin, Z.(2025) J Adv Res 68: 415-428
- PubMed: 38431124
- DOI: https://doi.org/10.1016/j.jare.2024.02.016
- Primary Citation Related Structures:
8JB4 - PubMed Abstract:
Antimicrobial peptides (AMPs) are valuable alternatives to traditional antibiotics, possess a variety of potent biological activities and exhibit immunomodulatory effects that alleviate difficult-to-treat infections. Clarifying the structure-activity relationships of AMPs can direct the synthesis of desirable peptide therapeutics. In this study, the lipopolysaccharide-binding domain (LBD) was identified through machine learning-guided directed evolution, which acts as a functional domain of the anti-lipopolysaccharide factor family of AMPs identified from Marsupenaeus japonicus. LBD A-D was identified as an output of this algorithm, in which the original LBD Mj sequence was the input, and the three-dimensional solution structure of LBD B was determined using nuclear magnetic resonance. Furthermore, our study involved a comprehensive series of experiments, including morphological studies and in vitro and in vivo antibacterial tests. The NMR solution structure showed that LBD B possesses a circular extended structure with a disulfide crosslink at the terminus and two 3 10 -helices and exhibits a broad antimicrobial spectrum. In addition, scanning electron microscopy (SEM) and transmission electron microscopy (TEM) showed that LBD B induced the formation of a cluster of bacteria wrapped in a flexible coating that ruptured and consequently killed the bacteria. Finally, coinjection of LBD B , Vibrio alginolyticus and Staphylococcus aureus in vivo improved the survival of M. japonicus, demonstrating the promising therapeutic role of LBD B for treating infectious disease. The findings of this study pave the way for the rational drug design of activity-enhanced peptide antibiotics.
- Center for Biological Science and Technology, Advanced Institute of Natural Sciences, Beijing Normal University, Zhuhai, Guangdong 519087, China.
Organizational Affiliation: