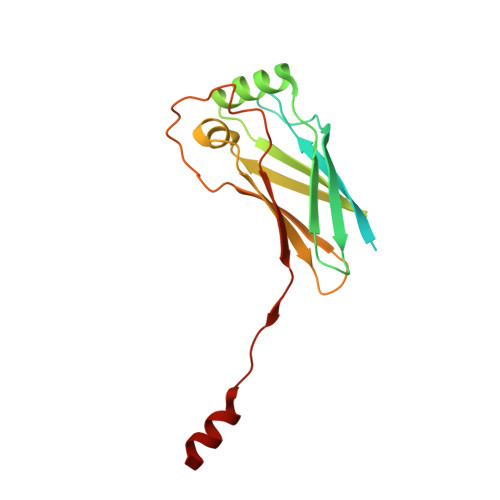
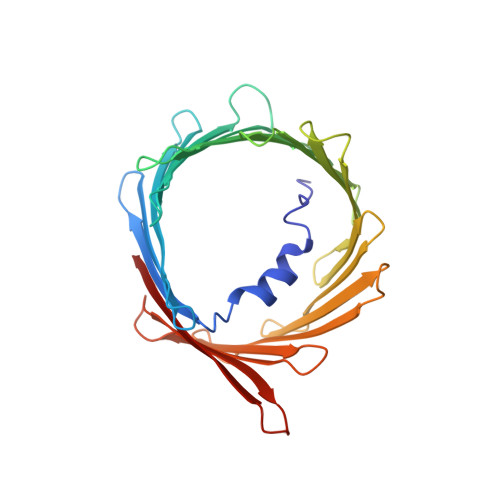
Structural insights into human EMC and its interaction with VDAC.
Li, M., Zhang, C., Xu, Y., Li, S., Huang, C., Wu, J., Lei, M.(2024) Aging (Albany NY) 16: 5501-5525
- PubMed: 38517390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.18632/aging.205660
- Primary Citation Related Structures:
8J0N, 8J0O - PubMed Abstract:
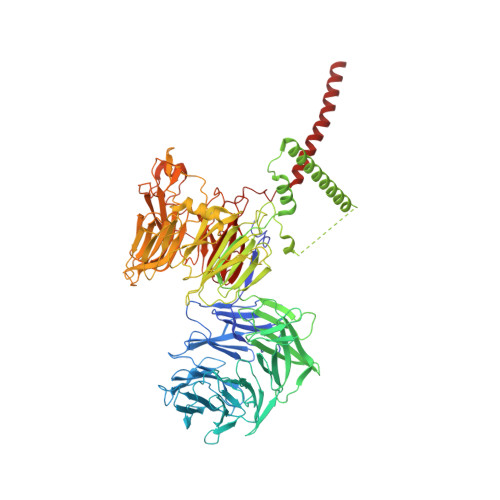
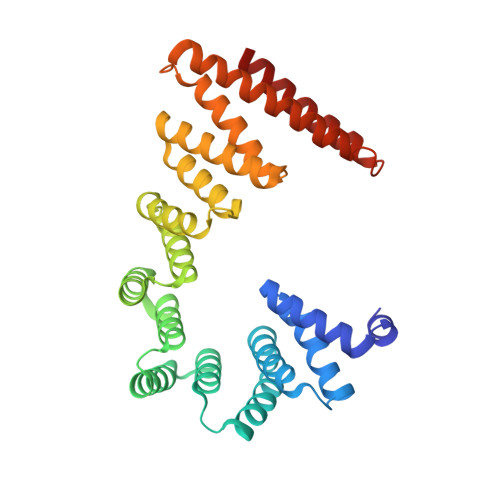
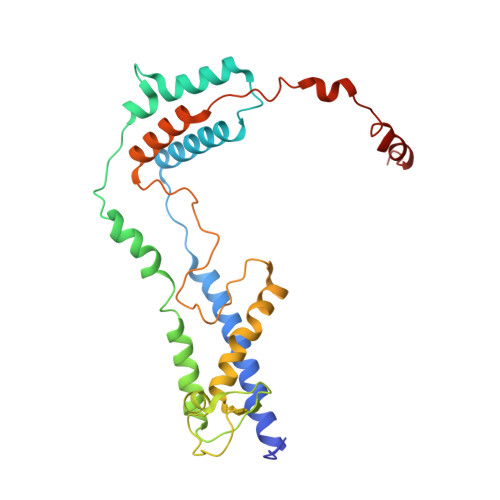
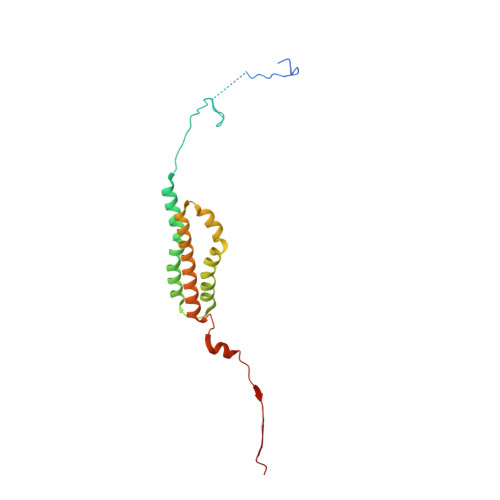
The endoplasmic reticulum (ER) membrane protein complex (EMC) is a conserved, multi-subunit complex acting as an insertase at the ER membrane. Growing evidence shows that the EMC is also involved in stabilizing and trafficking membrane proteins. However, the structural basis and regulation of its multifunctionality remain elusive. Here, we report cryo-electron microscopy structures of human EMC in apo- and voltage-dependent anion channel (VDAC)-bound states at resolutions of 3.47 Å and 3.32 Å, respectively. We discovered a specific interaction between VDAC proteins and the EMC at mitochondria-ER contact sites, which is conserved from yeast to humans. Moreover, we identified a gating plug located inside the EMC hydrophilic vestibule, the substrate-binding pocket for client insertion. Conformation changes of this gating plug during the apo-to-VDAC-bound transition reveal that the EMC unlikely acts as an insertase in the VDAC1-bound state. Based on the data analysis, the gating plug may regulate EMC functions by modifying the hydrophilic vestibule in different states. Our discovery offers valuable insights into the structural basis of EMC's multifunctionality.
- Ninth People’s Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai 200011, China.
Organizational Affiliation: