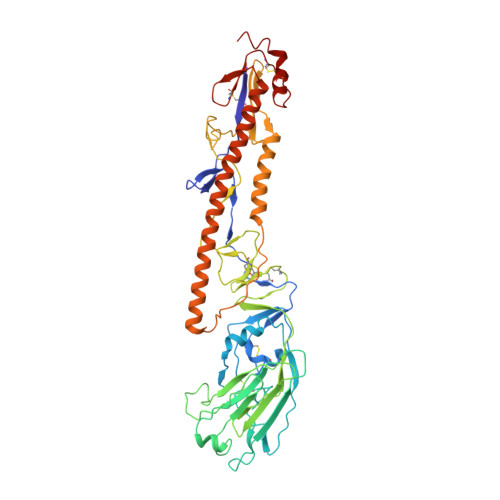
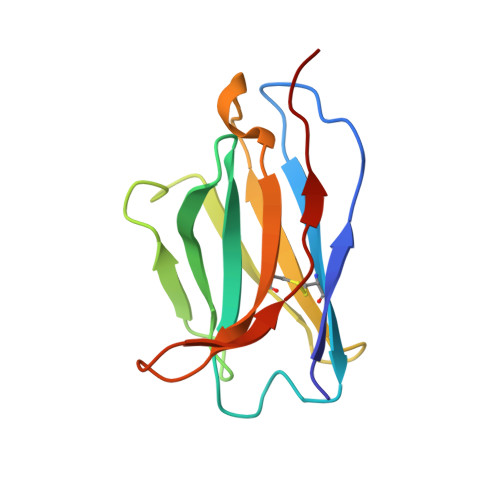
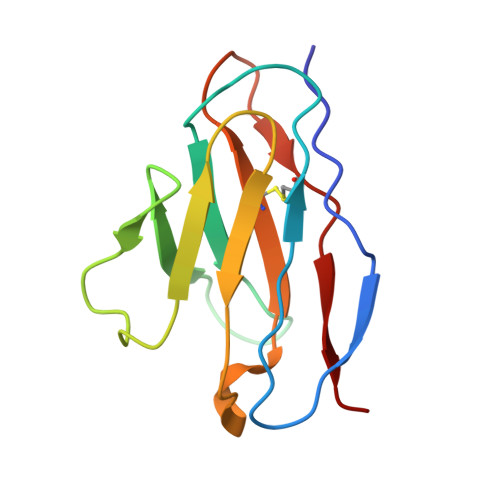
Structural basis of different neutralization capabilities of monoclonal antibodies against H7N9 virus.
Zhao, B., Sun, Z., Wang, S., Shi, Z., Jiang, Y., Wang, X., Deng, G., Jiao, P., Chen, H., Wang, J.(2025) J Virol 99: e0140024-e0140024
- PubMed: 39704525 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/jvi.01400-24
- Primary Citation Related Structures:
8IUX, 8IUY, 8IUZ, 8IV0 - PubMed Abstract:
Neutralizing antibodies (nAbs) are important for the treatment of emerging viral diseases and for effective vaccine development. In this study, we generated and evaluated three nAbs (1H9, 2D7, and C4H4) against H7N9 influenza viruses and found that they differ in their ability to inhibit viral attachment, membrane fusion, and egress. We resolved the cryo-electron microscopy (cryo-EM) structures of H7N9 hemagglutinin (HA) alone and in complex with the nAb antigen-binding fragments (Fabs) and identified the HA head-located epitope for each nAb, thereby revealing the molecular basis and key residues that determine the differences in these nAbs in neutralizing H7N9 viruses. Moreover, we found that the humanized nAb CC4H4 provided complete protection in mice against death caused by a lethal H7N9 virus infection, even when nAb was given 3 days after the mice were infected. These findings provide new insights into the neutralizing mechanism and structural basis for the rational design of H7N9 virus vaccines and therapeutics.IMPORTANCEH7N9 viruses have caused severe infections in both birds and humans since their emergence in early 2013 in China. Their persistent presence and variation in avian populations pose a significant threat to both poultry and humans. There are no treatments for human infections. In this study, we thoroughly investigated the neutralization mechanisms, structural basis, and therapeutic effects of three nAbs (1H9, 2D7, and C4H4) against H7N9 viruses. We revealed the molecular determinants underlying the varied performances of the three nAbs in neutralizing H7N9 viruses and protecting H7N9-infected mice. These insights provide a solid foundation for the rational design of vaccines and therapeutics against H7N9 viruses.
- State Key Laboratory for Animal Disease Control and Prevention & National Data Center for Animal Infectious Diseases, Harbin Veterinary Research Institute, Chinese Academy of Agricultural Sciences, Harbin, People's Republic of China.
Organizational Affiliation: