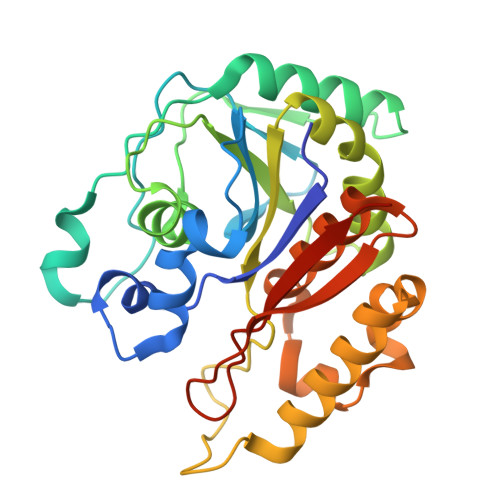
Structural Insights into 4,5-DOPA Extradiol Dioxygenase from Beta vulgaris : Unraveling the Key Step in Versatile Betalain Biosynthesis.
Chiang, C.C., Lu, Y.J., Liu, J.W., Lin, S.W., Chou, C.C., Lin, C.H., Chien, I.W., Hsu, C.H.(2025) J Agric Food Chem 73: 6785-6794
- PubMed: 40055856 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jafc.4c09501
- Primary Citation Related Structures:
8IN2 - PubMed Abstract:
Betalains, a group of pigments widely distributed in various plants, are extensively applied in the food, beverage, and medicinal industries. The biosynthesis of betalains involves the enzymatic action of 4,5-DOPA-dioxygenase, which catalyzes the key ring-opening reaction of DOPA to produce betalamic acid, a crucial intermediate in the pathway. The crystal structure of a 4,5-DOPA-dioxygenase from Beta vulgaris (BvDOD) was determined in this study. The structural analysis revealed that BvDOD exhibited a structural fold similar to that of other members of the extradiol dioxygenase family. Moreover, the Fe-ligand residues His15, His53, and His229 indicated the enzyme's reliance on nonheme iron for catalyzing the ring-opening reaction. Molecular docking and mutational analysis identified two conserved residues, His119 and His175, in the active site essential for the catalytic reaction. In addition, Thr17, Asp254, and Tyr260 contributed to properly positioning the substrate in the active site. This study has provided structural insights into substrate recognition and catalytic mechanisms of BvDOD, which can be applied to develop enzymes for improved betalain production.
- Department of Agricultural Chemistry, National Taiwan University, Taipei 10617, Taiwan.
Organizational Affiliation: