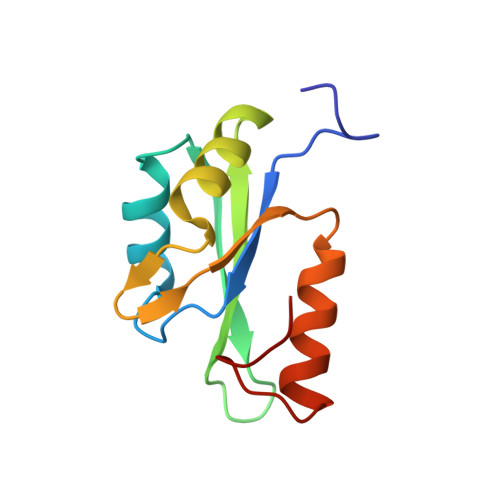
Molecular insight into the SETD1A/B N-terminal region and its interaction with WDR82.
Bao, S., Xu, C.(2023) Biochem Biophys Res Commun 658: 136-140
- PubMed: 37030068 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.03.064
- Primary Citation Related Structures:
8ILY, 8ILZ - PubMed Abstract:
SETD1A and SETD1B originate from Set1, the sole H3K4 methyltransferase in yeast, and they play important roles in active gene transcription. Here, we present the crystal structures of the RRM domains of human SETD1A and SETD1B. Although both RRM domains adopt a canonical RRM fold, their structural features are different from that of the yeast Set1 RRM domain, their yeast homolog. By using an ITC binding assay, we found an intrinsically disordered region in SETD1A/B binds WDR82. The structural analysis implies that the positively charged regions within human RRM domains might be involved in binding to RNA. Our work provides structural insight into the assembly of WDR82 with the catalytic subunits SETD1A/B in the context of the whole complex.
- MOE Key Laboratory for Membraneless Organelles & Cellular Dynamics, Hefei National Laboratory for Physical Sciences at the Microscale, School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, 230027, Hefei, PR China.
Organizational Affiliation: