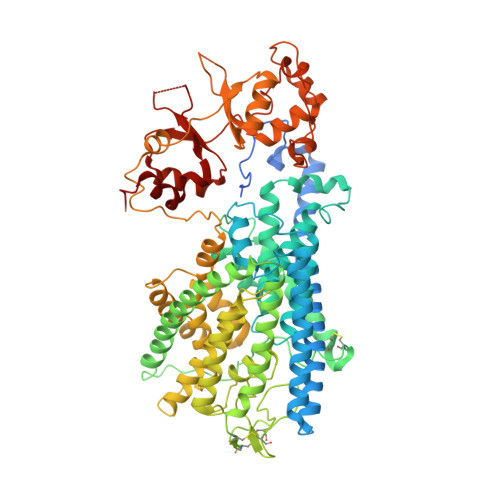
Molecular mechanism underlying regulation of Arabidopsis CLCa transporter by nucleotides and phospholipids.
Yang, Z., Zhang, X., Ye, S., Zheng, J., Huang, X., Yu, F., Chen, Z., Cai, S., Zhang, P.(2023) Nat Commun 14: 4879-4879
- PubMed: 37573431 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40624-z
- Primary Citation Related Structures:
8IAB, 8IAD - PubMed Abstract:
Chloride channels (CLCs) transport anion across membrane to regulate ion homeostasis and acidification of intracellular organelles, and are divided into anion channels and anion/proton antiporters. Arabidopsis thaliana CLCa (AtCLCa) transporter localizes to the tonoplast which imports NO 3 - and to a less extent Cl - from cytoplasm. The activity of AtCLCa and many other CLCs is regulated by nucleotides and phospholipids, however, the molecular mechanism remains unclear. Here we determine the cryo-EM structures of AtCLCa bound with NO 3 - and Cl - , respectively. Both structures are captured in ATP and PI(4,5)P 2 bound conformation. Structural and electrophysiological analyses reveal a previously unidentified N-terminal β-hairpin that is stabilized by ATP binding to block the anion transport pathway, thereby inhibiting the AtCLCa activity. While AMP loses the inhibition capacity due to lack of the β/γ- phosphates required for β-hairpin stabilization. This well explains how AtCLCa senses the ATP/AMP status to regulate the physiological nitrogen-carbon balance. Our data further show that PI(4,5)P 2 or PI(3,5)P 2 binds to the AtCLCa dimer interface and occupies the proton-exit pathway, which may help to understand the inhibition of AtCLCa by phospholipids to facilitate guard cell vacuole acidification and stomatal closure. In a word, our work suggests the regulatory mechanism of AtCLCa by nucleotides and phospholipids under certain physiological scenarios and provides new insights for future study of CLCs.
- National Key Laboratory of Plant Molecular Genetics, Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Chinese Academy of Sciences, Shanghai, 200032, China.
Organizational Affiliation: