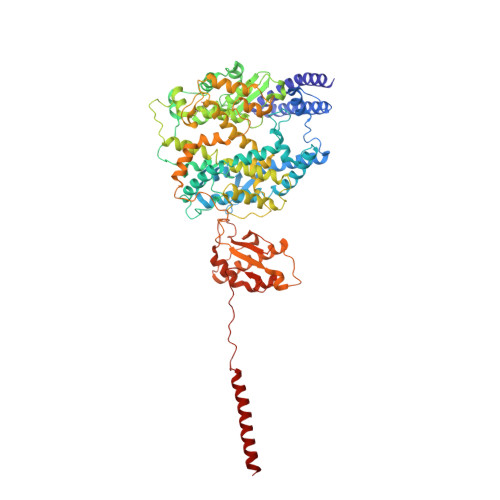
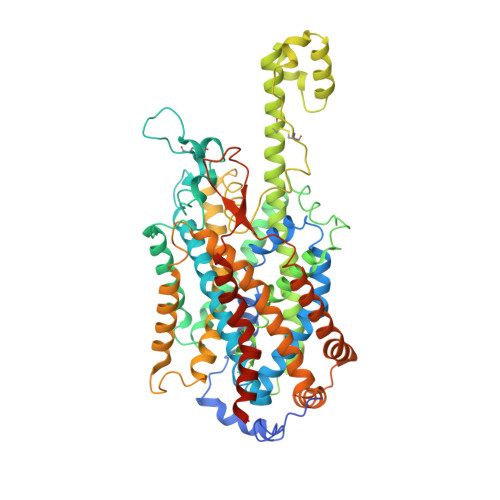
Structural insight into the substrate recognition and transport mechanism of amino acid transporter complex ACE2-B 0 AT1 and ACE2-SIT1.
Li, Y., Chen, Y., Zhang, Y., Shen, Y., Xu, K., Liu, Y., Wang, Z., Yan, R.(2023) Cell Discov 9: 93-93
- PubMed: 37684251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-023-00596-2
- Primary Citation Related Structures:
8I91, 8I92, 8I93 - Center for Infectious Disease Research, Westlake Laboratory of Life Sciences and Biomedicine, Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, Zhejiang, China.
Organizational Affiliation: