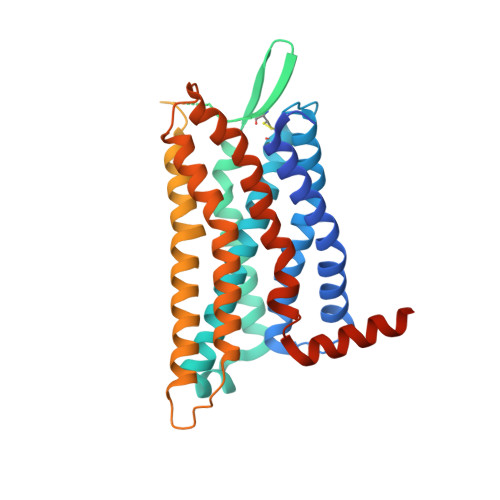
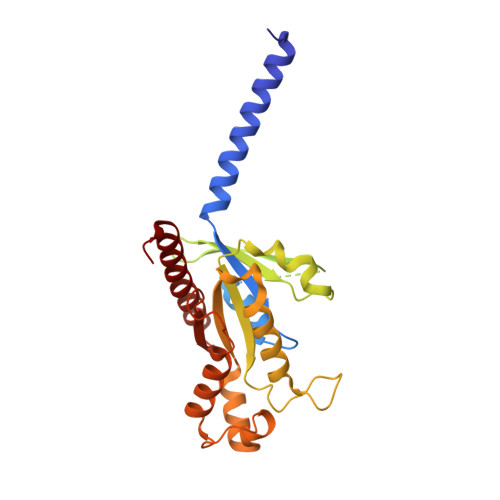
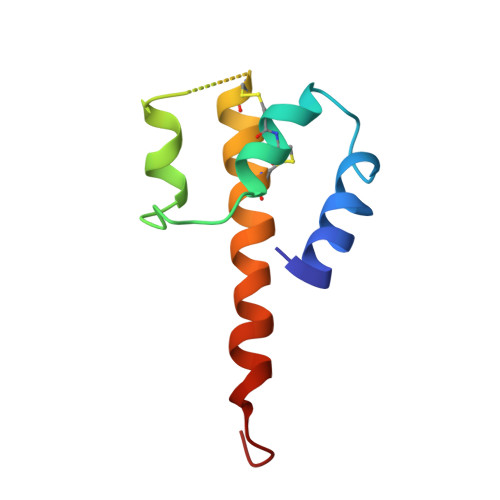
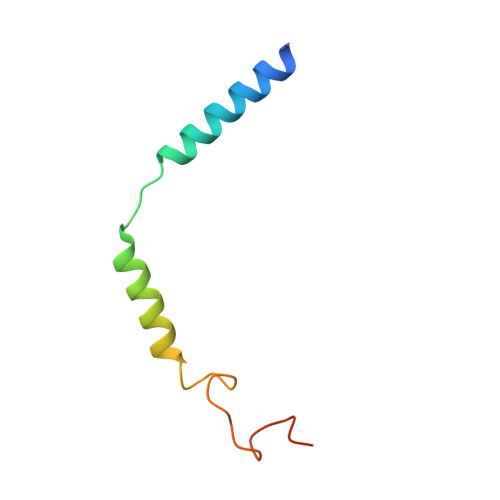
Revealing the signaling of complement receptors C3aR and C5aR1 by anaphylatoxins.
Wang, Y., Liu, W., Xu, Y., He, X., Yuan, Q., Luo, P., Fan, W., Zhu, J., Zhang, X., Cheng, X., Jiang, Y., Xu, H.E., Zhuang, Y.(2023) Nat Chem Biol 19: 1351-1360
- PubMed: 37169960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01339-w
- Primary Citation Related Structures:
8HK2, 8HK3, 8HK5 - PubMed Abstract:
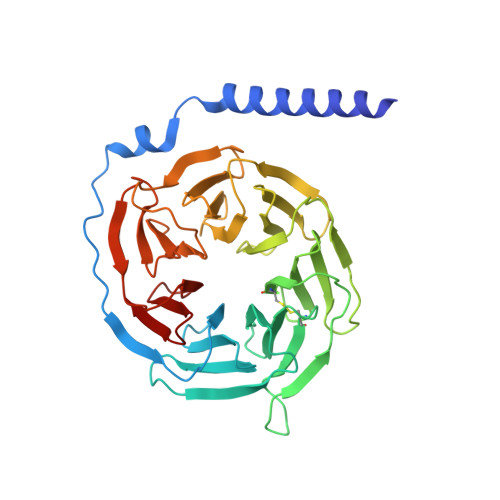
The complement receptors C3aR and C5aR1, whose signaling is selectively activated by anaphylatoxins C3a and C5a, are important regulators of both innate and adaptive immune responses. Dysregulations of C3aR and C5aR1 signaling lead to multiple inflammatory disorders, including sepsis, asthma and acute respiratory distress syndrome. The mechanism underlying endogenous anaphylatoxin recognition and activation of C3aR and C5aR1 remains elusive. Here we reported the structures of C3a-bound C3aR and C5a-bound C5aR1 as well as an apo-C3aR structure. These structures, combined with mutagenesis analysis, reveal a conserved recognition pattern of anaphylatoxins to the complement receptors that is different from chemokine receptors, unique pocket topologies of C3aR and C5aR1 that mediate ligand selectivity, and a common mechanism of receptor activation. These results provide crucial insights into the molecular understanding of C3aR and C5aR1 signaling and structural templates for rational drug design for treating inflammation disorders.
- State Key Laboratory of Drug Research, Center for Structure and Function of Drug Targets, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: