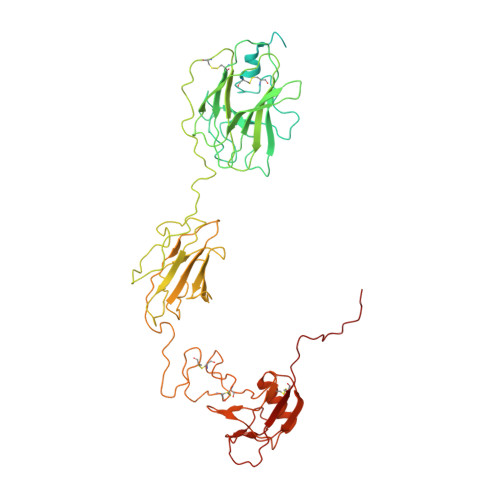
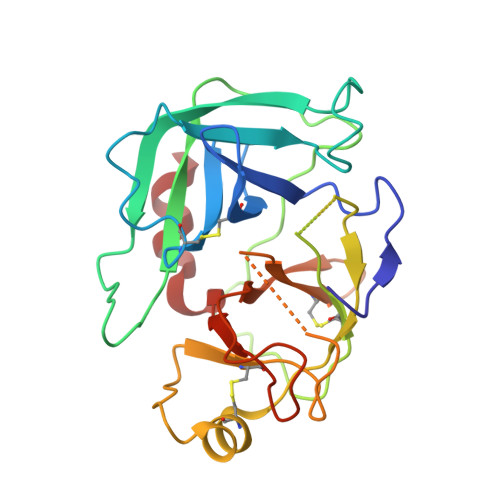
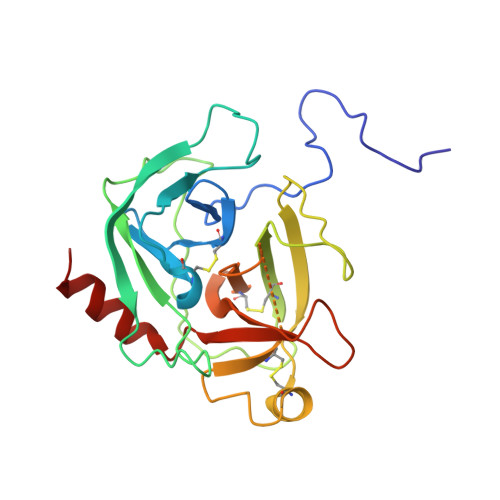
Cryo-EM structures reveal the activation and substrate recognition mechanism of human enteropeptidase.
Yang, X., Ding, Z., Peng, L., Song, Q., Zhang, D., Cui, F., Xia, C., Li, K., Yin, H., Li, S., Li, Z., Huang, H.(2022) Nat Commun 13: 6955-6955
- PubMed: 36376282
- DOI: https://doi.org/10.1038/s41467-022-34364-9
- Primary Citation Related Structures:
7WQW, 7WQX, 7WQZ, 7WR7, 8H3S, 8H3U - PubMed Abstract:
Enteropeptidase (EP) initiates intestinal digestion by proteolytically processing trypsinogen, generating catalytically active trypsin. EP dysfunction causes a series of pancreatic diseases including acute necrotizing pancreatitis. However, the molecular mechanisms of EP activation and substrate recognition remain elusive, due to the lack of structural information on the EP heavy chain. Here, we report cryo-EM structures of human EP in inactive, active, and substrate-bound states at resolutions from 2.7 to 4.9 Å. The EP heavy chain was observed to clamp the light chain with CUB2 domain for substrate recognition. The EP light chain N-terminus induced a rearrangement of surface-loops from inactive to active conformations, resulting in activated EP. The heavy chain then served as a hinge for light-chain conformational changes to recruit and subsequently cleave substrate. Our study provides structural insights into rearrangements of EP surface-loops and heavy chain dynamics in the EP catalytic cycle, advancing our understanding of EP-associated pancreatitis.
- Department of Gastroenterology, Changhai Hospital, Navy/Second Military Medical University, Shanghai, China.
Organizational Affiliation: