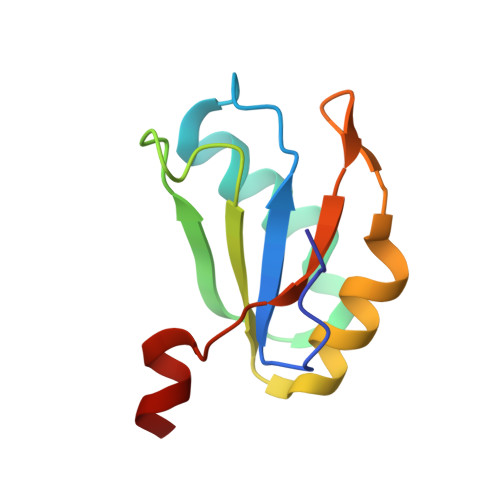
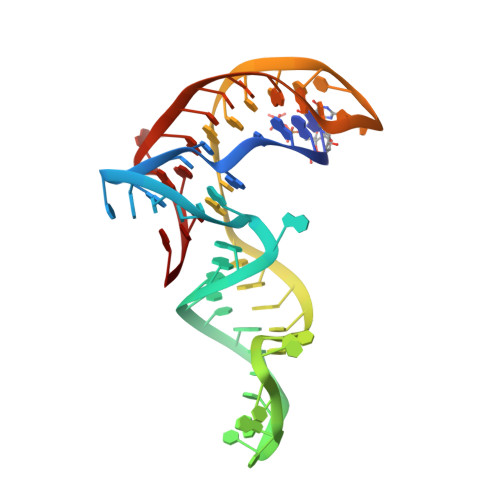
Structure-based investigations of the NAD+-II riboswitch.
Xu, X., Egger, M., Li, C., Chen, H., Micura, R., Ren, A.(2023) Nucleic Acids Res 51: 54-67
- PubMed: 36610789 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac1227
- Primary Citation Related Structures:
8GXB, 8GXC - PubMed Abstract:
Riboswitches are conserved non-coding domains in bacterial mRNA with gene regulation function that are essential for maintaining enzyme co-factor metabolism. Recently, the pnuC RNA motif was reported to selectively bind nicotinamide adenine dinucleotide (NAD+), defining a novel class of NAD+ riboswitches (NAD+-II) according to phylogenetic analysis. To reveal the three-dimensional architecture and the ligand-binding mode of this riboswitch, we solved the crystal structure of NAD+-II riboswitch in complex with NAD+. Strikingly and in contrast to class-I riboswitches that form a tight recognition pocket for the adenosine diphosphate (ADP) moiety of NAD+, the class-II riboswitches form a binding pocket for the nicotinamide mononucleotide (NMN) portion of NAD+ and display only unspecific interactions with the adenosine. We support this finding by an additional structure of the class-II RNA in complex with NMN alone. The structures define a novel RNA tertiary fold that was further confirmed by mutational analysis in combination with isothermal titration calorimetry (ITC), and 2-aminopurine-based fluorescence spectroscopic folding studies. Furthermore, we truncated the pnuC RNA motif to a short RNA helical scaffold with binding affinity comparable to the wild-type motif to allude to the potential of engineering the NAD+-II motif for biotechnological applications.
- Department of Gastroenterology/Department of Cardiology of the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, Zhejiang 310058, China.
Organizational Affiliation: