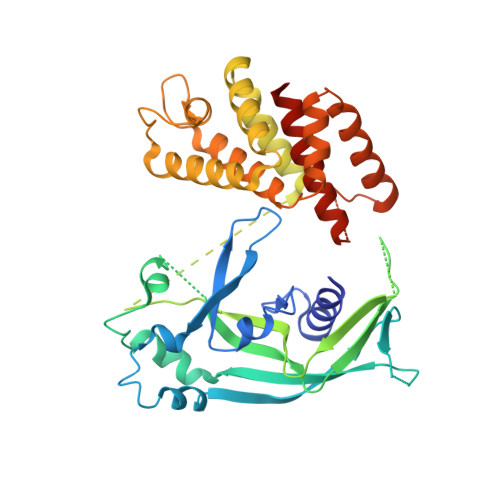
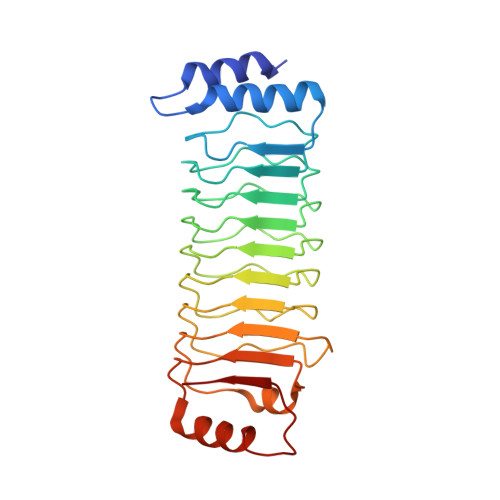
Structural mechanisms for regulation of GSDMB pore-forming activity.
Zhong, X., Zeng, H., Zhou, Z., Su, Y., Cheng, H., Hou, Y., She, Y., Feng, N., Wang, J., Shao, F., Ding, J.(2023) Nature 616: 598-605
- PubMed: 36991125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-023-05872-5
- Primary Citation Related Structures:
8GTJ, 8GTK, 8GTN - PubMed Abstract:
Cytotoxic lymphocyte-derived granzyme A (GZMA) cleaves GSDMB, a gasdermin-family pore-forming protein 1,2 , to trigger target cell pyroptosis 3 . GSDMB and the charter gasdermin family member GSDMD 4,5 have been inconsistently reported to be degraded by the Shigella flexneri ubiquitin-ligase virulence factor IpaH7.8 (refs. 6,7 ). Whether and how IpaH7.8 targets both gasdermins is undefined, and the pyroptosis function of GSDMB has even been questioned recently 6,8 . Here we report the crystal structure of the IpaH7.8-GSDMB complex, which shows how IpaH7.8 recognizes the GSDMB pore-forming domain. We clarify that IpaH7.8 targets human (but not mouse) GSDMD through a similar mechanism. The structure of full-length GSDMB suggests stronger autoinhibition than in other gasdermins 9,10 . GSDMB has multiple splicing isoforms that are equally targeted by IpaH7.8 but exhibit contrasting pyroptotic activities. Presence of exon 6 in the isoforms dictates the pore-forming, pyroptotic activity in GSDMB. We determine the cryo-electron microscopy structure of the 27-fold-symmetric GSDMB pore and depict conformational changes that drive pore formation. The structure uncovers an essential role for exon-6-derived elements in pore assembly, explaining pyroptosis deficiency in the non-canonical splicing isoform used in recent studies 6,8 . Different cancer cell lines have markedly different isoform compositions, correlating with the onset and extent of pyroptosis following GZMA stimulation. Our study illustrates fine regulation of GSDMB pore-forming activity by pathogenic bacteria and mRNA splicing and defines the underlying structural mechanisms.
- Peking University-Tsinghua University-National Institute of Biological Sciences Joint Graduate Program, School of Life Sciences, Tsinghua University, Beijing, PR China.
Organizational Affiliation: