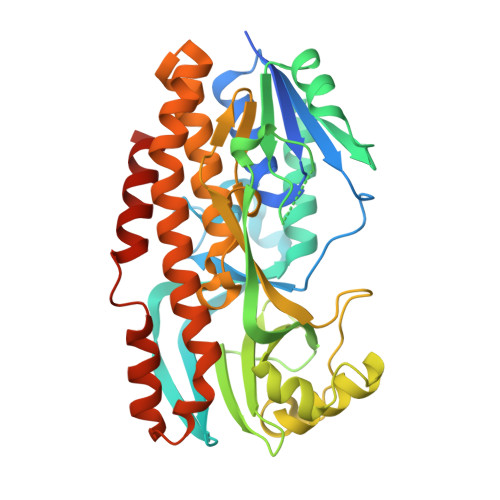
Structural basis for substrate binding and catalytic mechanism of the key enzyme VioD in the violacein synthesis pathway.
Xu, M., Xu, D., Gao, M., Zhuang, X., Wang, W., Sun, B., Ran, T.(2023) Proteins 91: 956-966
- PubMed: 36869636 Search on PubMed
- DOI: https://doi.org/10.1002/prot.26484
- Primary Citation Related Structures:
8GQR, 8H0M - PubMed Abstract:
Violacein is a pigment synthesized by gram-negative bacteria with various biological activities such as antimicrobial, antiviral, and anticancer activities. VioD is a key oxygenase converting protodeoxyviolaceinic acid to protoviolaceinic acid in violacein biosynthesis. To elucidate the catalytic mechanism of VioD, here, we resolved two crystal structures of VioD, a binary complex structure containing VioD and a FAD and a ternary complex structure composed of VioD, a FAD and a 2-ethyl-1-hexanol (EHN). Structural analysis revealed a deep funnel like binding pocket with wide entrance, this pocket is positively charged. The EHN is located at the deep bottom of the binding pocket near isoalloxazine ring. Further docking simulation help us to propose the mechanism of the hydroxylation of the substrate catalyzed by VioD. Bioinformatic analysis suggested and emphasized the importance of the conserved residues involved in substrate binding. Our results provide a structural basis for the catalytic mechanism of VioD.
- Department of Microbiology, College of Life Sciences, Nanjing Agricultural University, Nanjing, China.
Organizational Affiliation: