A conformation-specific nanobody targeting the nicotinamide mononucleotide-activated state of SARM1.
Hou, Y.N., Cai, Y., Li, W.H., He, W.M., Zhao, Z.Y., Zhu, W.J., Wang, Q., Mai, X., Liu, J., Lee, H.C., Stjepanovic, G., Zhang, H., Zhao, Y.J.(2022) Nat Commun 13: 7898-7898
- PubMed: 36550129 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-35581-y
- Primary Citation Related Structures:
8GNI, 8GNJ, 8GQ5 - PubMed Abstract:
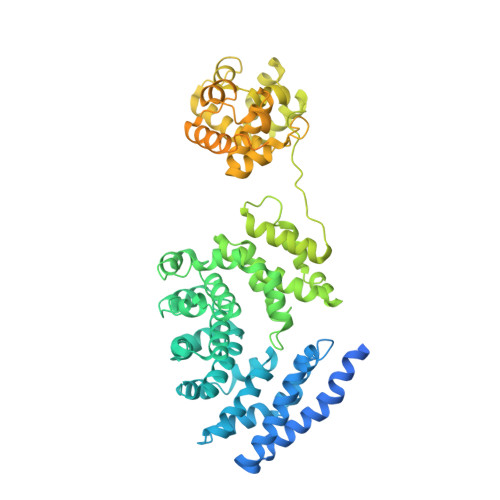
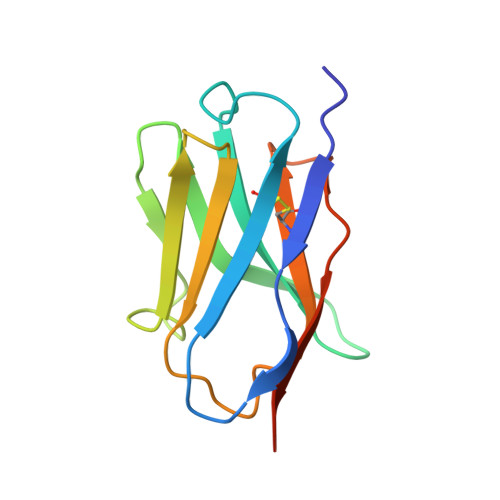
Sterile alpha (SAM) and Toll/interleukin-1 receptor (TIR) motif containing 1 (SARM1) is an autoinhibitory NAD-consuming enzyme that is activated by the accumulation of nicotinamide mononucleotide (NMN) during axonal injury. Its activation mechanism is not fully understood. Here, we generate a nanobody, Nb-C6, that specifically recognizes NMN-activated SARM1. Nb-C6 stains only the activated SARM1 in cells stimulated with CZ-48, a permeant mimetic of NMN, and partially activates SARM1 in vitro and in cells. Cryo-EM of NMN/SARM1/Nb-C6 complex shows an octameric structure with ARM domains bending significantly inward and swinging out together with TIR domains. Nb-C6 binds to SAM domain of the activated SARM1 and stabilized its ARM domain. Mass spectrometry analyses indicate that the activated SARM1 in solution is highly dynamic and that the neighboring TIRs form transient dimers via the surface close to one BB loop. We show that Nb-C6 is a valuable tool for studies of SARM1 activation.
- State Key Laboratory of Chemical Oncogenomics, Key Laboratory of Chemical Genomics, Peking University Shenzhen Graduate School, Shenzhen, 518055, China.
Organizational Affiliation: