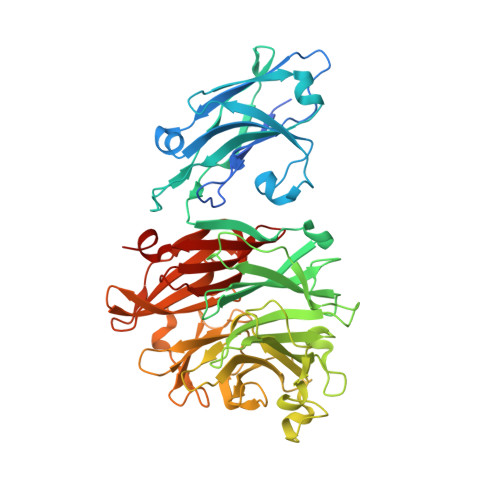
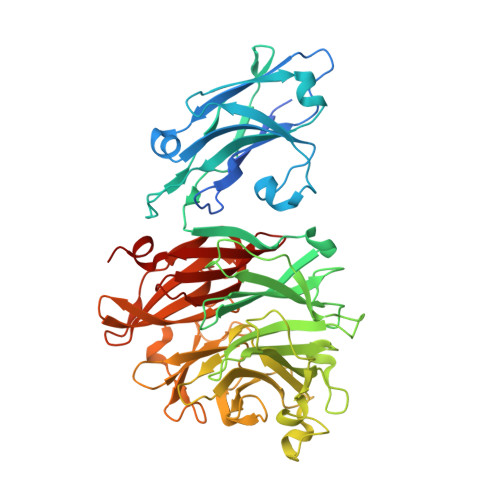
Structural and enzymatic characterization of the sialidase SiaPG from Porphyromonas gingivalis.
Dong, W.B., Jiang, Y.L., Zhu, Z.L., Zhu, J., Li, Y., Xia, R., Zhou, K.(2023) Acta Crystallogr F Struct Biol Commun 79: 87-94
- PubMed: 36995120 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23001735
- Primary Citation Related Structures:
8GN6 - PubMed Abstract:
The sialidases, which catalyze the hydrolysis of sialic acid from extracellular glycoconjugates, are a group of major virulence factors in various pathogenic bacteria. In Porphyromonas gingivalis, which causes human periodontal disease, sialidase contributes to bacterial pathogenesis via promoting the formation of biofilms and capsules, reducing the ability for macrophage clearance, and providing nutrients for bacterial colonization. Here, the crystal structure of the P. gingivalis sialidase SiaPG is reported at 2.1 Å resolution, revealing an N-terminal carbohydrate-binding domain followed by a canonical C-terminal catalytic domain. Simulation of the product sialic acid in the active-site pocket together with functional analysis enables clear identification of the key residues that are required for substrate binding and catalysis. Moreover, structural comparison with other sialidases reveals distinct features of the active-site pocket which might confer substrate specificity. These findings provide the structural basis for the further design and optimization of effective inhibitors to target SiaPG to fight against P. gingivalis-derived oral diseases.
- Department of Stomatology, The Second Affiliated Hospital of Anhui Medical University, Hefei, Anhui 230601, People's Republic of China.
Organizational Affiliation: