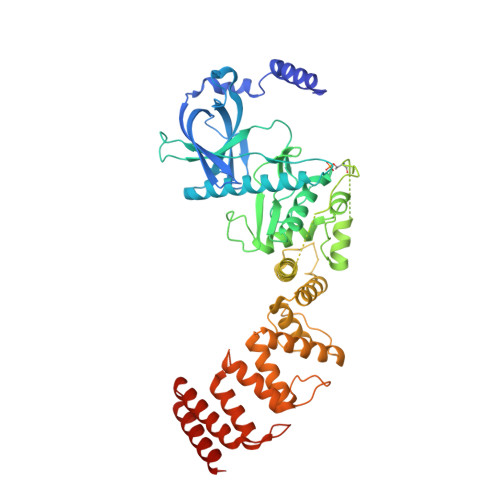
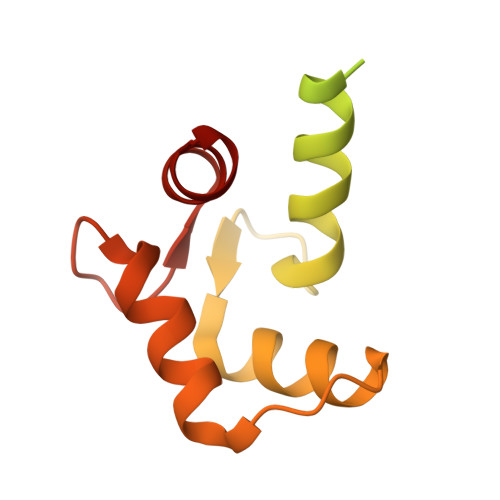
Structure of the complex between calmodulin and a functional construct of eukaryotic elongation factor 2 kinase bound to an ATP-competitive inhibitor.
Piserchio, A., Isiorho, E.A., Dalby, K.N., Ghose, R.(2023) J Biological Chem 299: 104813-104813
- PubMed: 37172726 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.104813
- Primary Citation Related Structures:
8GM4, 8GM5 - PubMed Abstract:
The calmodulin-activated α-kinase, eukaryotic elongation factor 2 kinase (eEF-2K), serves as a master regulator of translational elongation by specifically phosphorylating and reducing the ribosome affinity of the guanosine triphosphatase, eukaryotic elongation factor 2 (eEF-2). Given its critical role in a fundamental cellular process, dysregulation of eEF-2K has been implicated in several human diseases, including those of the cardiovascular system, chronic neuropathies, and many cancers, making it a critical pharmacological target. In the absence of high-resolution structural information, high-throughput screening efforts have yielded small-molecule candidates that show promise as eEF-2K antagonists. Principal among these is the ATP-competitive pyrido-pyrimidinedione inhibitor, A-484954, which shows high specificity toward eEF-2K relative to a panel of "typical" protein kinases. A-484954 has been shown to have some degree of efficacy in animal models of several disease states. It has also been widely deployed as a reagent in eEF-2K-specific biochemical and cell-biological studies. However, given the absence of structural information, the precise mechanism of the A-484954-mediated inhibition of eEF-2K has remained obscure. Leveraging our identification of the calmodulin-activatable catalytic core of eEF-2K, and our recent determination of its long-elusive structure, here we present the structural basis for its specific inhibition by A-484954. This structure, which represents the first for an inhibitor-bound catalytic domain of a member of the α-kinase family, enables rationalization of the existing structure-activity relationship data for A-484954 variants and lays the groundwork for further optimization of this scaffold to attain enhanced specificity/potency against eEF-2K.
- Department of Chemistry and Biochemistry, The City College of New York, New York, New York, USA.
Organizational Affiliation: