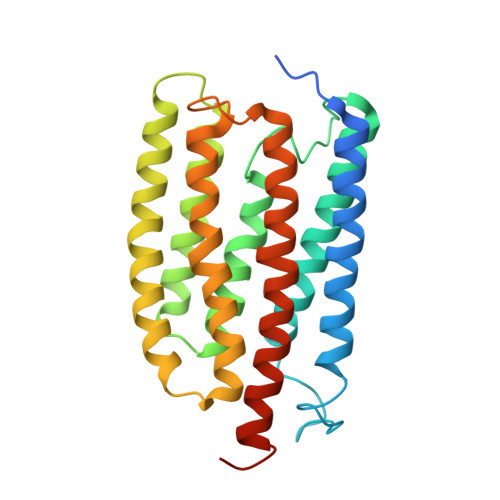
Structures of channelrhodopsin paralogs in peptidiscs explain their contrasting K + and Na + selectivities.
Morizumi, T., Kim, K., Li, H., Govorunova, E.G., Sineshchekov, O.A., Wang, Y., Zheng, L., Bertalan, E., Bondar, A.N., Askari, A., Brown, L.S., Spudich, J.L., Ernst, O.P.(2023) Nat Commun 14: 4365-4365
- PubMed: 37474513 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40041-2
- Primary Citation Related Structures:
8GI8, 8GI9 - PubMed Abstract:
Kalium channelrhodopsin 1 from Hyphochytrium catenoides (HcKCR1) is a light-gated channel used for optogenetic silencing of mammalian neurons. It selects K + over Na + in the absence of the canonical tetrameric K + selectivity filter found universally in voltage- and ligand-gated channels. The genome of H. catenoides also encodes a highly homologous cation channelrhodopsin (HcCCR), a Na + channel with >100-fold larger Na + to K + permeability ratio. Here, we use cryo-electron microscopy to determine atomic structures of these two channels embedded in peptidiscs to elucidate structural foundations of their dramatically different cation selectivity. Together with structure-guided mutagenesis, we show that K + versus Na + selectivity is determined at two distinct sites on the putative ion conduction pathway: in a patch of critical residues in the intracellular segment (Leu69/Phe69, Ile73/Ser73 and Asp116) and within a cluster of aromatic residues in the extracellular segment (primarily, Trp102 and Tyr222). The two filters are on the opposite sides of the photoactive site involved in channel gating.
- Department of Biochemistry, University of Toronto, Toronto, ON, Canada.
Organizational Affiliation: