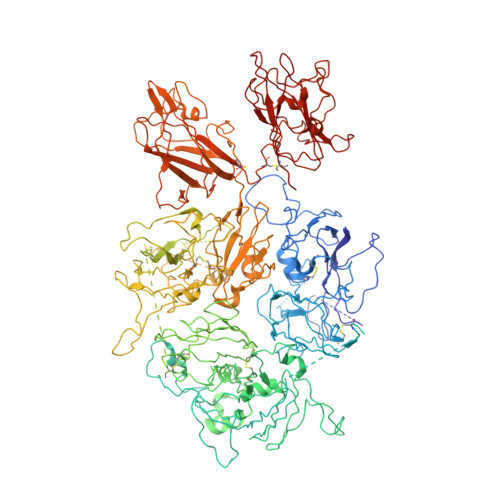
Structure of coagulation factor VIII bound to a patient-derived anti-C1 domain antibody inhibitor.
Childers, K.C., Avery, N.G., Estrada Alamo, K.A., Davulcu, O., Haynes, R.M., Lollar, P., Doering, C.B., Coxon, C.H., Spiegel, P.C.(2023) Blood 142: 197-201
- PubMed: 37192299 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood.2023020181
- Primary Citation Related Structures:
8G6I - PubMed Abstract:
The development of pathogenic antibody inhibitors against coagulation factor VIII (FVIII) occurs in ∼30% of patients with congenital hemophilia A receiving FVIII replacement therapy, as well as in all cases of acquired hemophilia A. KM33 is an anti-C1 domain antibody inhibitor previously isolated from a patient with severe hemophilia A. In addition to potently blocking FVIII binding to von Willebrand factor and phospholipid surfaces, KM33 disrupts FVIII binding to lipoprotein receptor-related protein 1 (LRP1), which drives FVIII hepatic clearance and antigen presentation in dendritic cells. Here, we report on the structure of FVIII bound to NB33, a recombinant derivative of KM33, via single-particle cryo-electron microscopy. Structural analysis revealed that the NB33 epitope localizes to the FVIII residues R2090-S2094 and I2158-R2159, which constitute membrane-binding loops in the C1 domain. Further analysis revealed that multiple FVIII lysine and arginine residues, previously shown to mediate binding to LRP1, dock onto an acidic cleft at the NB33 variable domain interface, thus blocking a putative LRP1 binding site. Together, these results demonstrate a novel mechanism of FVIII inhibition by a patient-derived antibody inhibitor and provide structural evidence for engineering FVIII with reduced LRP1-mediated clearance.
- Chemistry Department, Western Washington University, Bellingham, WA.
Organizational Affiliation: