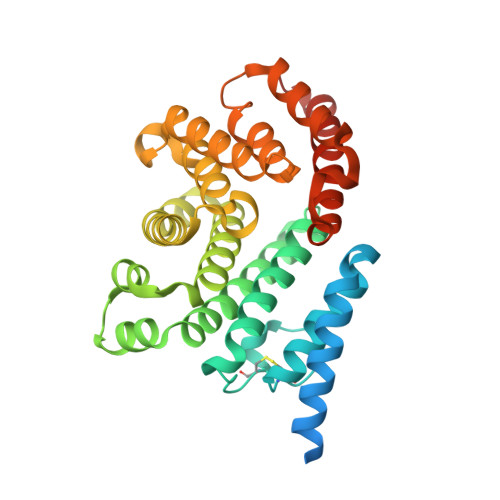
Structures and Mechanisms of a Novel Bacterial Transport System for Fatty Acids.
Zhai, L., Chou, J.C., Oo, H., Dassama, L.M.K.(2023) Chembiochem 24: e202300156-e202300156
- PubMed: 37170829 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.202300156
- Primary Citation Related Structures:
8G52, 8G53 - PubMed Abstract:
Bacterial acquisition of metabolites is largely facilitated by transporters with unique substrate scopes. The tripartite ATP-independent periplasmic (TRAP) transporters comprise a large family of bacterial proteins that facilitate the uptake of a variety of small molecules. It has been reported that some TRAP systems encode a fourth protein, the T component. The T-component, or TatT, is predicted to be a periplasmic-facing lipoprotein that enables the uptake of metabolites from the outer membrane. However, no substrates were revealed for any TatT and their functional role(s) remained enigmatic. We recently identified a homolog in Methylococcus capsulatus that binds to sterols, and herein, we report two additional homologs that demonstrate a preference for long-chain fatty acids. Our bioinformatics, quantitative analyses of protein-ligand interactions, and high-resolution crystal structures suggest that TatTs might facilitate the trafficking of hydrophobic or lipophilic substrates and represent a new class of bacterial lipid and fatty acid transporters.
- Department of Chemistry and Sarafan ChEM-H Institute, Stanford University, Stanford, CA, 94305, USA.
Organizational Affiliation: