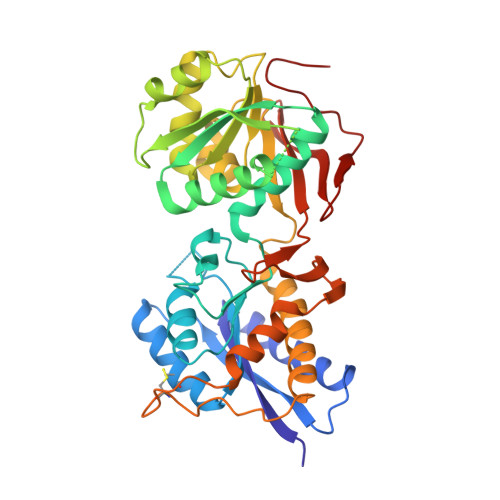
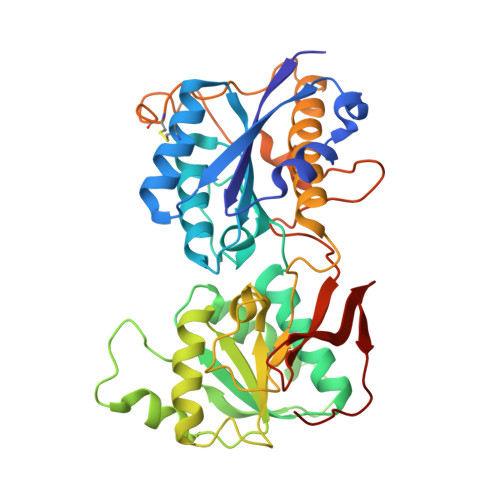
Novel GluN2B-Selective NMDA Receptor Negative Allosteric Modulator Possesses Intrinsic Analgesic Properties and Enhances Analgesia of Morphine in a Rodent Tail Flick Pain Model.
Harris, L.D., Regan, M.C., Myers, S.J., Nocilla, K.A., Akins, N.S., Tahirovic, Y.A., Wilson, L.J., Dingledine, R., Furukawa, H., Traynelis, S.F., Liotta, D.C.(2023) ACS Chem Neurosci 14: 917-935
- PubMed: 36779874 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschemneuro.2c00779
- Primary Citation Related Structures:
8G18 - PubMed Abstract:
Many cases of accidental death associated with drug overdose are due to chronic opioid use, tolerance, and addiction. Analgesic tolerance is characterized by a decreased response to the analgesic effects of opioids, requiring increasingly higher doses to maintain the desired level of pain relief. Overactivation of GluN2B-containing N -methyl-d-Aspartate receptors is thought to play a key role in mechanisms underlying cellular adaptation that takes place in the development of analgesic tolerance. Herein, we describe a novel GluN2B-selective negative allosteric modulator, EU93-108 , that shows high potency and brain penetrance. We describe the structural basis for binding at atomic resolution. This compound possesses intrinsic analgesic properties in the rodent tail immersion test. EU93-108 has an acute and significant anodyne effect, whereby morphine when combined with EU93-108 produces a higher tail flick latency compared to that of morphine alone. These data suggest that engagement of GluN2B as a target has utility in the treatment of pain, and EU93-108 could serve as an appropriate tool compound to interrogate this hypothesis. Future structure-activity relationship work around this scaffold could give rise to compounds that can be co-administered with opioids to diminish the onset of tolerance due to chronic opioid use, thereby modifying their utility.
- Department of Chemistry, Emory University, Atlanta, Georgia30322, United States.
Organizational Affiliation: