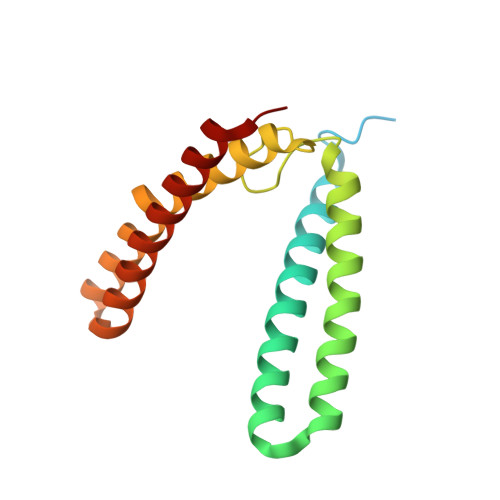
Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Budziszewski, G.R., Lynch, M.L., Snell, M.E., Monteiro, D.C., Bowman, S.E.(2025) bioRxiv
- PubMed: 40501712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.06.01.657255
- Primary Citation Related Structures:
8FUH, 8FVV, 8FXD, 9ONM, 9ONN, 9ONO, 9ONQ, 9ONR - PubMed Abstract:
Rubrerythrins are a group of proteins within the Ferritin-like superfamily that display a defining four-helix bundle domain. They also show multiple structural features that are crucial to their functionality as iron storage proteins and in detoxification and oxidative stress response. Here we investigate rubrerythrin (Rbr) in multiple metalated states, from the pathogen Burkholderia pseudomallei ( Bp ). We use X-ray crystallography for structure determination of Rbr to probe the capacity and specificity of metal binding. Bp Rbr lacks the rubredoxin moiety found in canonical Rbrs from anaerobic lineages, and we demonstrate that Bp Rbr also possesses a domain-swapped dimer, which has functional implications for its putative role in oxidative stress response. We also carry out in crystallo spectroscopic assessment of Bp Rbr with various metals, using energy dispersive X-ray (EDX) spectroscopy. We observe that samples can contain metals other than those supplied in crystallization conditions, and developed a strategy of utilizing EDX spectroscopy to select those samples with single metal incorporation for downstream diffraction data collection. Our work underscores the importance of spectroscopic probing for definitive metal identification and characterization. Present structures of apo-, Fe-, Mn-, and Co-bound rubrerythrin from B. pseudomallei Bp Rbr lacks rubredoxin domain and possesses unique domain swap fold in the dimer Bp Rbr features include promiscuous metal binding and pre-formed metal binding sites Highlights importance of in crystallo spectroscopy in investigating metalloproteins Mutant Rbr with abrogated metal binding supports premise of pre-formed binding sites.