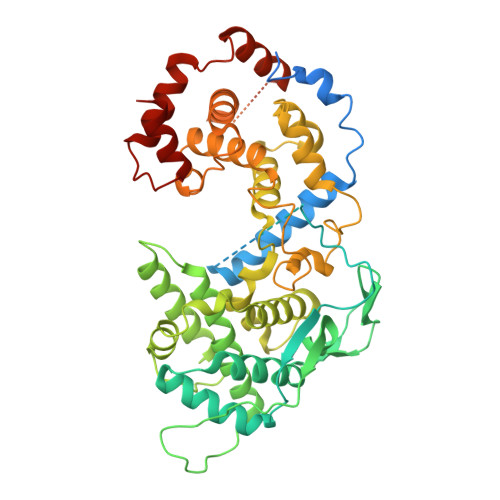
Structural Determination of the Australian Bat Lyssavirus Nucleoprotein and Phosphoprotein Complex.
Donnelly, C.M., Stewart, M., Roby, J.A., Sundaramoorthy, V., Forwood, J.K.(2023) Viruses 16: 33-33
- PubMed: 38229694 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/v16010033
- Primary Citation Related Structures:
8FWL - PubMed Abstract:
Australian bat lyssavirus (ABLV) shows similar clinical symptoms as rabies, but there are currently no protein structures available for ABLV proteins. In lyssaviruses, the interaction between nucleoprotein (N) and phosphoprotein (N) in the absence of RNA generates a complex (N 0 P) that is crucial for viral assembly, and understanding the interface between these two proteins has the potential to provide insight into a key feature: the viral lifecycle. In this study, we used recombinant chimeric protein expression and X-ray crystallography to determine the structure of ABLV nucleoprotein bound to residues 1-40 of its phosphoprotein chaperone. Comparison of our results with the recently generated structure of RABV CVS-11 N 0 P demonstrated a highly conserved interface in this complex. Because the N 0 P interface is conserved in the lyssaviruses of phylogroup I, it is an attractive therapeutic target for multiple rabies-causing viral species.
- School of Dentistry and Medical Sciences, Charles Sturt University, Wagga Wagga, NSW 2678, Australia.
Organizational Affiliation: