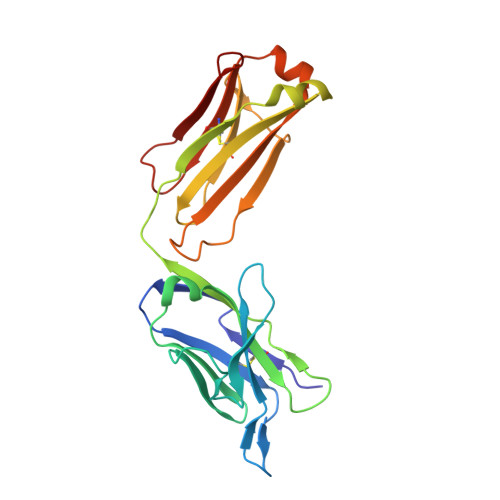
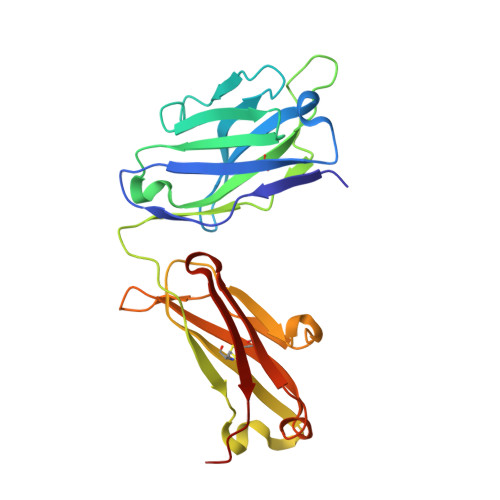
Inadequate structural constraint on Fab approach rather than paratope elicitation limits HIV-1 MPER vaccine utility.
Tan, K., Chen, J., Kaku, Y., Wang, Y., Donius, L., Khan, R.A., Li, X., Richter, H., Seaman, M.S., Walz, T., Hwang, W., Reinherz, E.L., Kim, M.(2023) Nat Commun 14: 7218-7218
- PubMed: 37940661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-42097-6
- Primary Citation Related Structures:
8FWF, 8FXJ, 8FYM, 8FZ2 - PubMed Abstract:
Broadly neutralizing antibodies (bnAbs) against HIV-1 target conserved envelope (Env) epitopes to block viral replication. Here, using structural analyses, we provide evidence to explain why a vaccine targeting the membrane-proximal external region (MPER) of HIV-1 elicits antibodies with human bnAb-like paratopes paradoxically unable to bind HIV-1. Unlike in natural infection, vaccination with MPER/liposomes lacks a necessary structure-based constraint to select for antibodies with an adequate approach angle. Consequently, the resulting Abs cannot physically access the MPER crawlspace on the virion surface. By studying naturally arising Abs, we further reveal that flexibility of the human IgG3 hinge mitigates the epitope inaccessibility and additionally facilitates Env spike protein crosslinking. Our results suggest that generation of IgG3 subtype class-switched B cells is a strategy for anti-MPER bnAb induction. Moreover, the findings illustrate the need to incorporate topological features of the target epitope in immunogen design.
- Structural Biology Center, X-ray Science Division, Advanced Photon Source, Argonne National Laboratory, Lemont, IL, USA.
Organizational Affiliation: