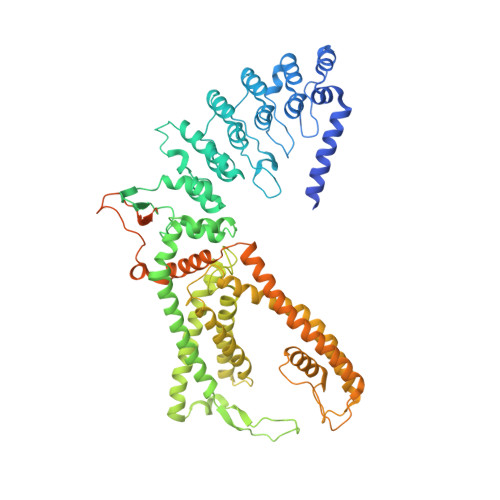
Structural mechanism of human oncochannel TRPV6 inhibition by the natural phytoestrogen genistein.
Neuberger, A., Trofimov, Y.A., Yelshanskaya, M.V., Nadezhdin, K.D., Krylov, N.A., Efremov, R.G., Sobolevsky, A.I.(2023) Nat Commun 14: 2659-2659
- PubMed: 37160865 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38352-5
- Primary Citation Related Structures:
8FOA, 8FOB - PubMed Abstract:
Calcium-selective oncochannel TRPV6 is the major driver of cell proliferation in human cancers. While significant effort has been invested in the development of synthetic TRPV6 inhibitors, natural channel blockers have been largely neglected. Here we report the structure of human TRPV6 in complex with the plant-derived phytoestrogen genistein, extracted from Styphnolobium japonicum, that was shown to inhibit cell invasion and metastasis in cancer clinical trials. Despite the pharmacological value, the molecular mechanism of TRPV6 inhibition by genistein has remained enigmatic. We use cryo-EM combined with electrophysiology, calcium imaging, mutagenesis, and molecular dynamics simulations to show that genistein binds in the intracellular half of the TRPV6 pore and acts as an ion channel blocker and gating modifier. Genistein binding to the open channel causes pore closure and a two-fold symmetrical conformational rearrangement in the S4-S5 and S6-TRP helix regions. The unprecedented mechanism of TRPV6 inhibition by genistein uncovers new possibilities in structure-based drug design.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY, USA.
Organizational Affiliation: