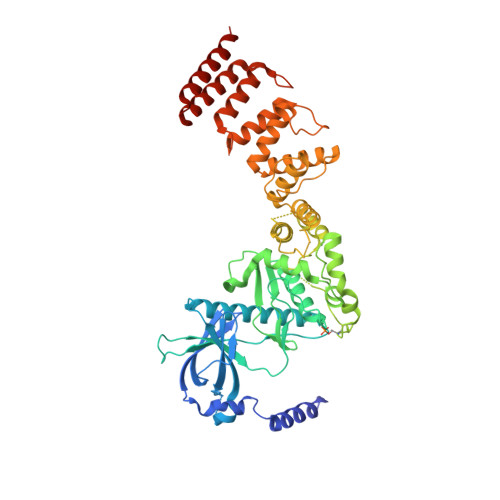
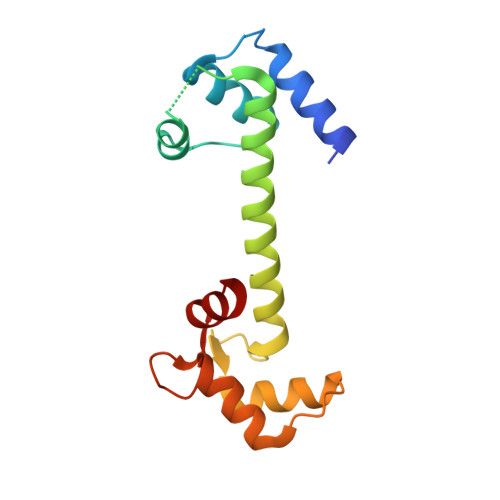
ADP enhances the allosteric activation of eukaryotic elongation factor 2 kinase by calmodulin.
Piserchio, A., Long, K.J., Browning, L.S., Bohanon, A.L., Isiorho, E.A., Dalby, K.N., Ghose, R.(2023) Proc Natl Acad Sci U S A 120: e2300902120-e2300902120
- PubMed: 37068230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2300902120
- Primary Citation Related Structures:
8FNY, 8FO6 - PubMed Abstract:
Protein translation, one of the most energy-consumptive processes in a eukaryotic cell, requires robust regulation, especially under energy-deprived conditions. A critical component of this regulation is the suppression of translational elongation through reduced ribosome association of the GTPase eukaryotic elongation factor 2 (eEF-2) resulting from its specific phosphorylation by the calmodulin (CaM)-activated α-kinase eEF-2 kinase (eEF-2K). It has been suggested that the eEF-2K response to reduced cellular energy levels is indirect and mediated by the universal energy sensor AMP-activated protein kinase (AMPK) through direct stimulatory phosphorylation and/or downregulation of the eEF-2K-inhibitory nutrient-sensing mTOR pathway. Here, we provide structural, biochemical, and cell-biological evidence of a direct energy-sensing role of eEF-2K through its stimulation by ADP. A crystal structure of the nucleotide-bound complex between CaM and the functional core of eEF-2K phosphorylated at its primary stimulatory site (T348) reveals ADP bound at a unique pocket located on the face opposite that housing the kinase active site. Within this basic pocket (BP), created at the CaM/eEF-2K interface upon complex formation, ADP is stabilized through numerous interactions with both interacting partners. Biochemical analyses using wild-type eEF-2K and specific BP mutants indicate that ADP stabilizes CaM within the active complex, increasing the sensitivity of the kinase to CaM. Induction of energy stress through glycolysis inhibition results in significantly reduced enhancement of phosphorylated eEF-2 levels in cells expressing ADP-binding compromised BP mutants compared to cells expressing wild-type eEF-2K. These results suggest a direct energy-sensing role for eEF-2K through its cooperative interaction with CaM and ADP.
- Department of Chemistry and Biochemistry, The City College of New York, New York, NY 10031.
Organizational Affiliation: