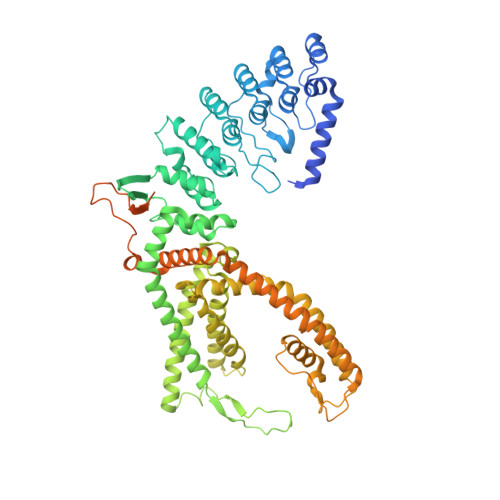
Structural basis of the activation of TRPV5 channels by long-chain acyl-Coenzyme-A.
Lee, B.H., De Jesus Perez, J.J., Moiseenkova-Bell, V., Rohacs, T.(2023) Nat Commun 14: 5883-5883
- PubMed: 37735536 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-41577-z
- Primary Citation Related Structures:
8FFO, 8FHH, 8FHI - PubMed Abstract:
Long-chain acyl-coenzyme A (LC-CoA) is a crucial metabolic intermediate that plays important cellular regulatory roles, including activation and inhibition of ion channels. The structural basis of ion channel regulation by LC-CoA is not known. Transient receptor potential vanilloid 5 and 6 (TRPV5 and TRPV6) are epithelial calcium-selective ion channels. Here, we demonstrate that LC-CoA activates TRPV5 and TRPV6 in inside-out patches, and both exogenously supplied and endogenously produced LC-CoA can substitute for the natural ligand phosphatidylinositol 4,5-bisphosphate (PI(4,5)P 2 ) in maintaining channel activity in intact cells. Utilizing cryo-electron microscopy, we determined the structure of LC-CoA-bound TRPV5, revealing an open configuration with LC-CoA occupying the same binding site as PI(4,5)P 2 in previous studies. This is consistent with our finding that PI(4,5)P 2 could not further activate the channels in the presence of LC-CoA. Our data provide molecular insights into ion channel regulation by a metabolic signaling molecule.
- Department of Pharmacology, Physiology and Neuroscience, Rutgers, New Jersey Medical School, Newark, NJ, USA.
Organizational Affiliation: