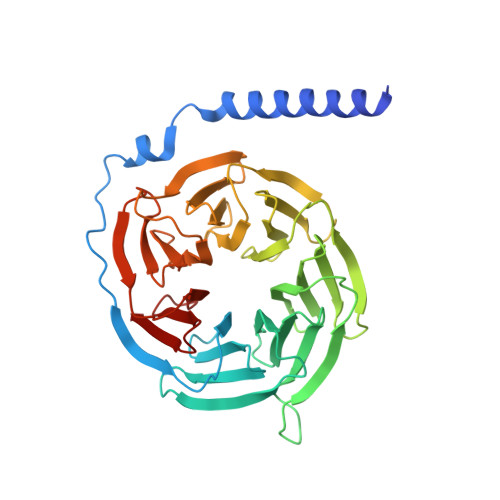
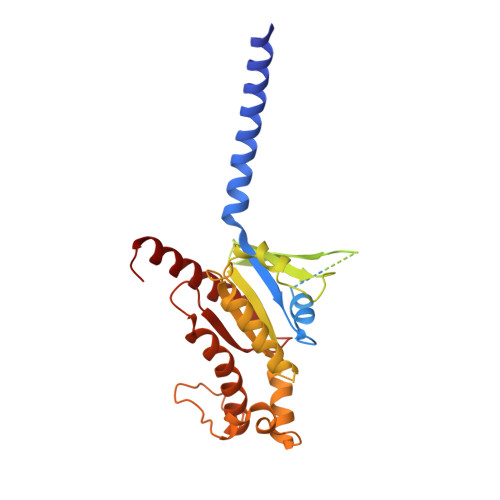
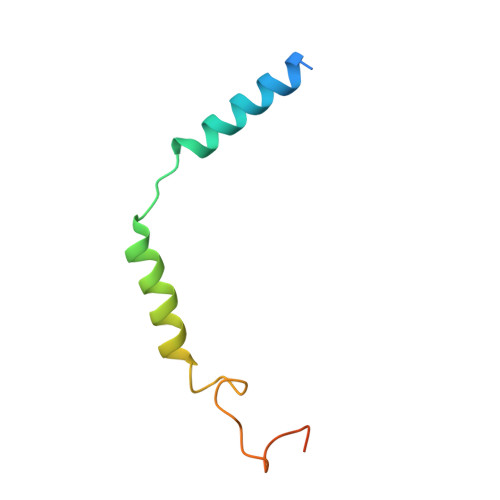
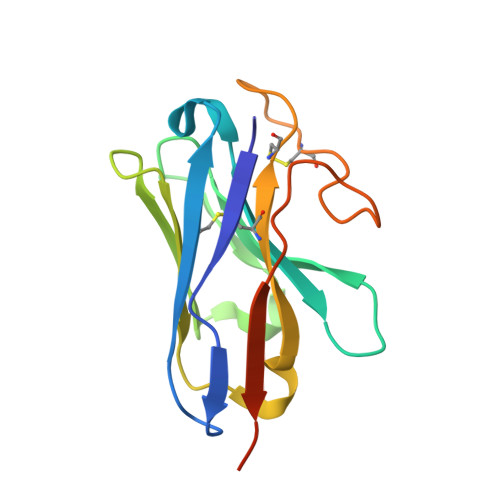
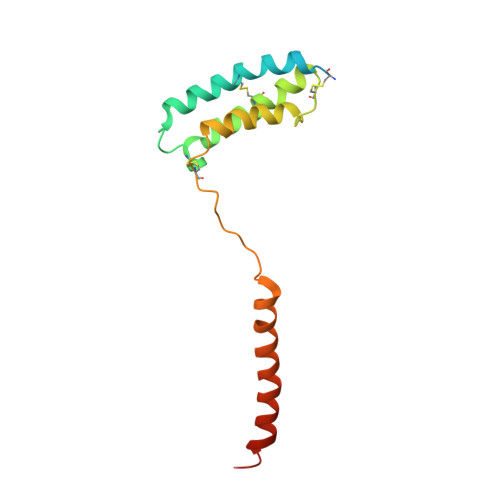
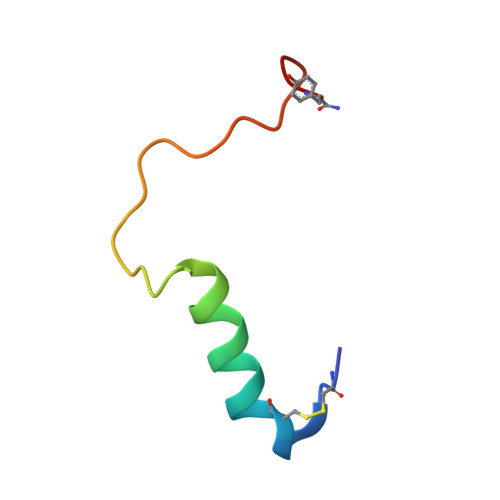
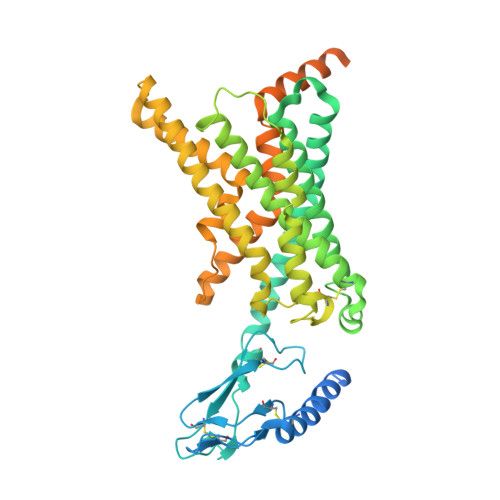
Structural insight into selectivity of amylin and calcitonin receptor agonists.
Cao, J., Belousoff, M.J., Gerrard, E., Danev, R., Fletcher, M.M., Dal Maso, E., Schreuder, H., Lorenz, K., Evers, A., Tiwari, G., Besenius, M., Li, Z., Johnson, R.M., Wootten, D., Sexton, P.M.(2024) Nat Chem Biol 20: 162-169
- PubMed: 37537379 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01393-4
- Primary Citation Related Structures:
8F0J, 8F0K, 8F2A, 8F2B - PubMed Abstract:
Amylin receptors (AMYRs), heterodimers of the calcitonin receptor (CTR) and one of three receptor activity-modifying proteins, are promising obesity targets. A hallmark of AMYR activation by Amy is the formation of a 'bypass' secondary structural motif (residues S19-P25). This study explored potential tuning of peptide selectivity through modification to residues 19-22, resulting in a selective AMYR agonist, San385, as well as nonselective dual amylin and calcitonin receptor agonists (DACRAs), with San45 being an exemplar. We determined the structure and dynamics of San385-bound AMY 3 R, and San45 bound to AMY 3 R or CTR. San45, via its conjugated lipid at position 21, was anchored at the edge of the receptor bundle, enabling a stable, alternative binding mode when bound to the CTR, in addition to the bypass mode of binding to AMY 3 R. Targeted lipid modification may provide a single intervention strategy for design of long-acting, nonselective, Amy-based DACRAs with potential anti-obesity effects.
- Drug Discovery Biology Theme, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, Victoria, Australia.
Organizational Affiliation: