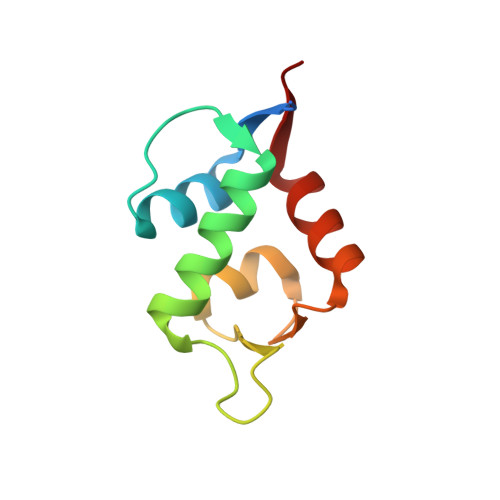
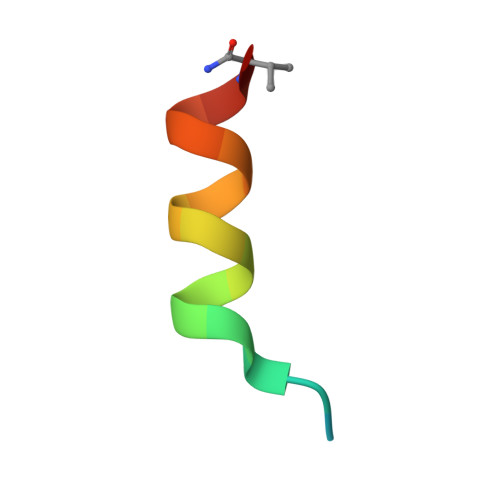
Single-Shot Flow Synthesis of D-Proteins for Mirror-Image Phage Display
Callahan, A.J., Gandhesiri, S., Travaline, T.L., Lozano Salazar, L., Hanna, S., Lee, Y.-C., Li, K., Tokareva, O.S., Swiecicki, J.-M., Loas, A., Verdine, G.L., McGee, J.H., Pentelute, B.L.(2023) ChemRxiv