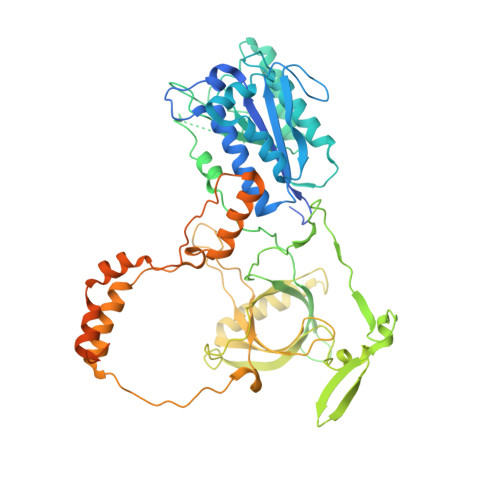
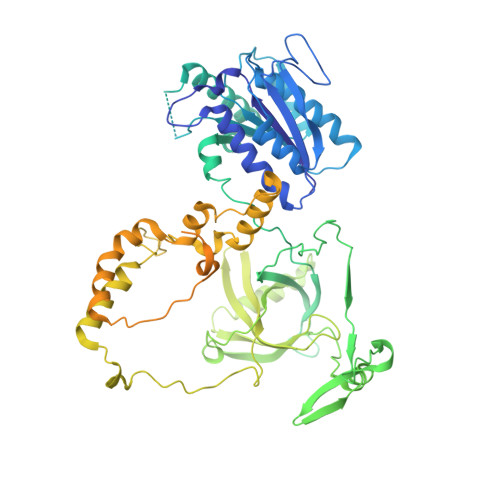
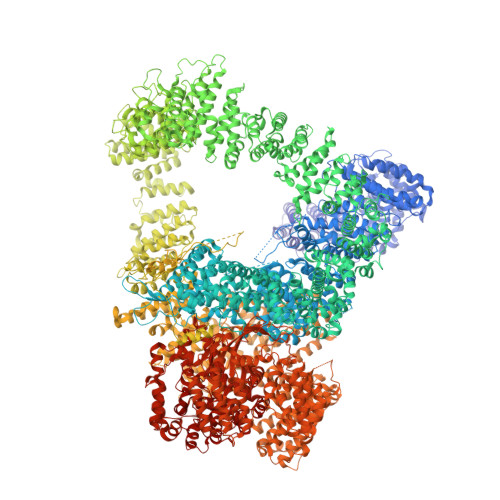
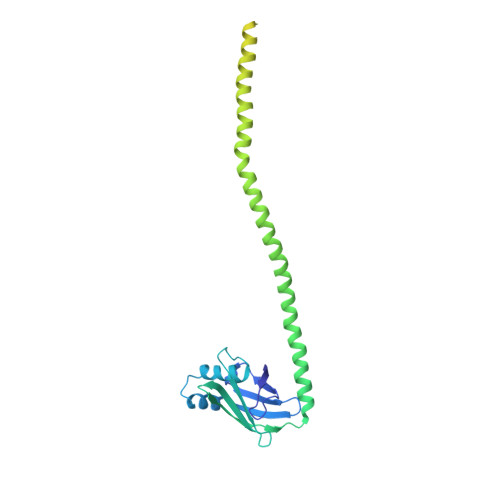
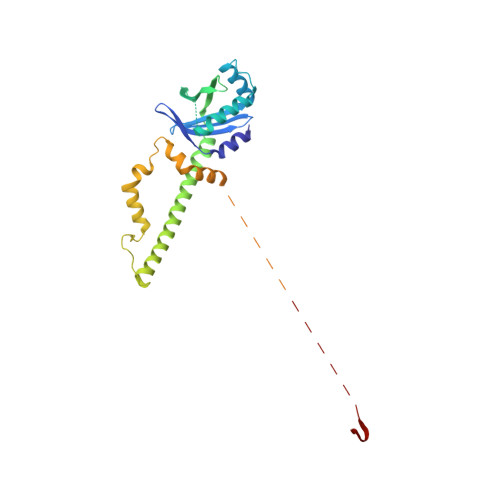
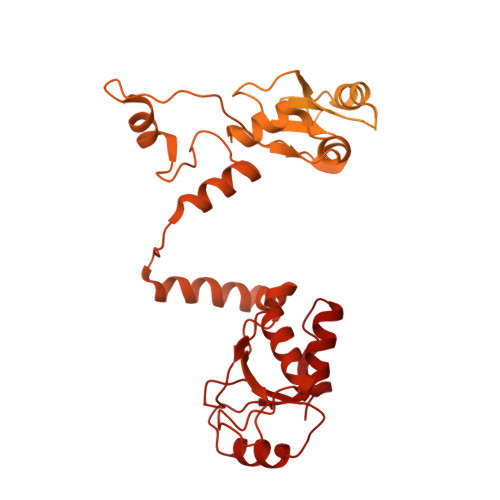
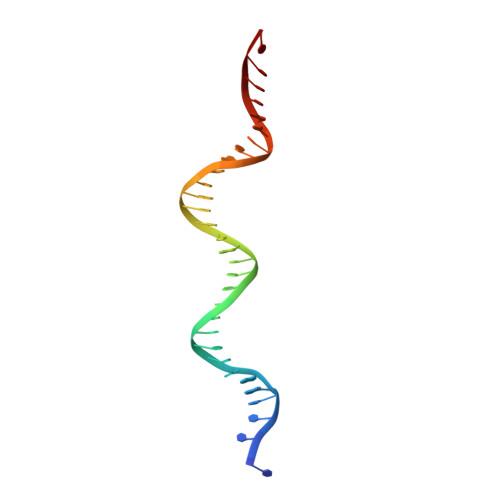
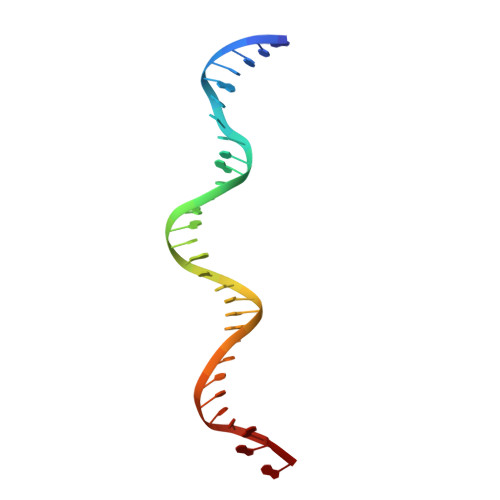
Cryo-EM visualization of DNA-PKcs structural intermediates in NHEJ.
Chen, S., Vogt, A., Lee, L., Naila, T., McKeown, R., Tomkinson, A.E., Lees-Miller, S.P., He, Y.(2023) Sci Adv 9: eadg2838-eadg2838
- PubMed: 37256947 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adg2838
- Primary Citation Related Structures:
8EZ9, 8EZA, 8EZB - PubMed Abstract:
DNA double-strand breaks (DSBs), one of the most cytotoxic forms of DNA damage, can be repaired by the tightly regulated nonhomologous end joining (NHEJ) machinery (Stinson and Loparo and Zhao et al. ). Core NHEJ factors form an initial long-range (LR) synaptic complex that transitions into a DNA-PKcs (DNA-dependent protein kinase, catalytic subunit)-free, short-range state to align the DSB ends (Chen et al. ). Using single-particle cryo-electron microscopy, we have visualized three additional key NHEJ complexes representing different transition states, with DNA-PKcs adopting distinct dimeric conformations within each of them. Upon DNA-PKcs autophosphorylation, the LR complex undergoes a substantial conformational change, with both Ku and DNA-PKcs rotating outward to promote DNA break exposure and DNA-PKcs dissociation. We also captured a dimeric state of catalytically inactive DNA-PKcs, which resembles structures of other PIKK (Phosphatidylinositol 3-kinase-related kinase) family kinases, revealing a model of the full regulatory cycle of DNA-PKcs during NHEJ.
- Department of Molecular Biosciences, Northwestern University, Evanston, IL, USA.
Organizational Affiliation: