Antibody-directed evolution reveals a mechanism for enhanced neutralization at the HIV-1 fusion peptide site.
Banach, B.B., Pletnev, S., Olia, A.S., Xu, K., Zhang, B., Rawi, R., Bylund, T., Doria-Rose, N.A., Nguyen, T.D., Fahad, A.S., Lee, M., Lin, B.C., Liu, T., Louder, M.K., Madan, B., McKee, K., O'Dell, S., Sastry, M., Schon, A., Bui, N., Shen, C.H., Wolfe, J.R., Chuang, G.Y., Mascola, J.R., Kwong, P.D., DeKosky, B.J.(2023) Nat Commun 14: 7593-7593
- PubMed: 37989731 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-42098-5
- Primary Citation Related Structures:
8ELI, 8EUU, 8EUV, 8EUW, 8F7Z - PubMed Abstract:
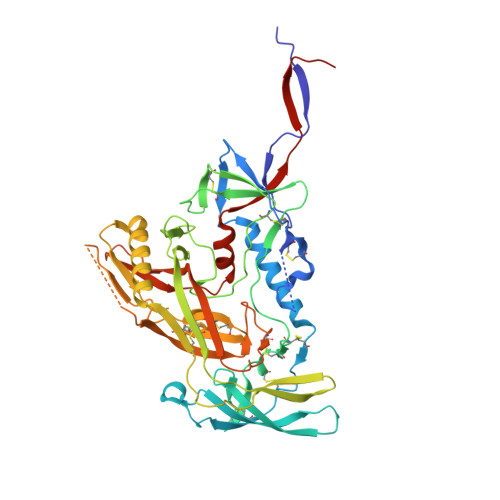
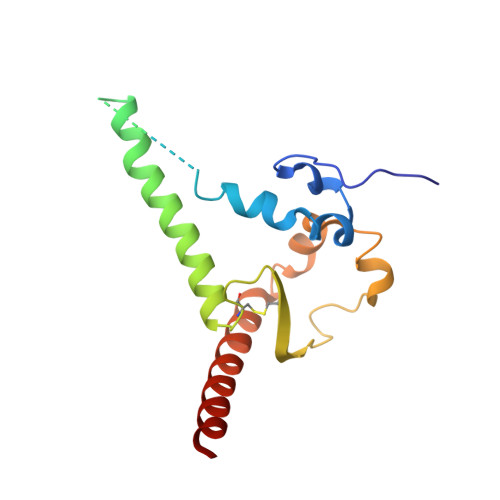
The HIV-1 fusion peptide (FP) represents a promising vaccine target, but global FP sequence diversity among circulating strains has limited anti-FP antibodies to ~60% neutralization breadth. Here we evolve the FP-targeting antibody VRC34.01 in vitro to enhance FP-neutralization using site saturation mutagenesis and yeast display. Successive rounds of directed evolution by iterative selection of antibodies for binding to resistant HIV-1 strains establish a variant, VRC34.01_mm28, as a best-in-class antibody with 10-fold enhanced potency compared to the template antibody and ~80% breadth on a cross-clade 208-strain neutralization panel. Structural analyses demonstrate that the improved paratope expands the FP binding groove to accommodate diverse FP sequences of different lengths while also recognizing the HIV-1 Env backbone. These data reveal critical antibody features for enhanced neutralization breadth and potency against the FP site of vulnerability and accelerate clinical development of broad HIV-1 FP-targeting vaccines and therapeutics.
- Bioengineering Graduate Program, The University of Kansas, Lawrence, KS, 66045, USA.
Organizational Affiliation: