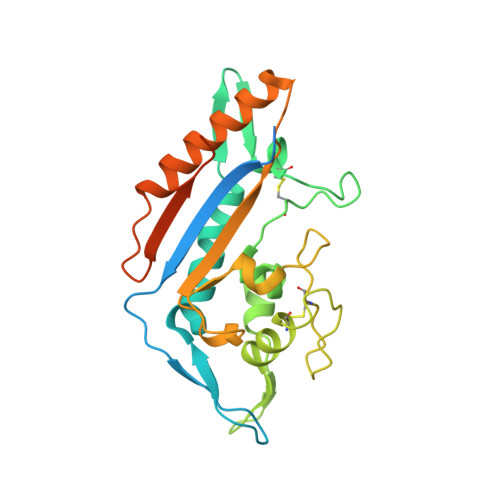
Purification and structure of luminal domain C of human Niemann-Pick C1 protein.
Odongo, L., Zadrozny, K.K., Diehl, W.E., Luban, J., White, J.M., Ganser-Pornillos, B.K., Tamm, L.K., Pornillos, O.(2023) Acta Crystallogr F Struct Biol Commun 79: 45-50
- PubMed: 36748341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23000705
- Primary Citation Related Structures:
8EUS - PubMed Abstract:
Niemann-Pick C1 protein (NPC1) is a membrane protein that primarily resides in late endosomes and lysosomes, and plays an important role in cholesterol homeostasis in the cell. The second luminal domain of NPC1 (NPC1-C) serves as the intracellular receptor for Ebola and Marburg viruses. Here, the recombinant production of nonglycosylated and glycosylated NPC1-C and a new crystal form of the nonglycosylated protein are reported. The crystals belonged to space group P2 1 and diffracted to 2.3 Å resolution. The structure is similar to other reported structures of NPC1-C, with differences observed in the protruding loops when compared with NPC1-C in complex with Ebola virus glycoprotein or NPC2.
- Center for Membrane and Cell Physiology, University of Virginia, Charlottesville, VA 22908, USA.
Organizational Affiliation: