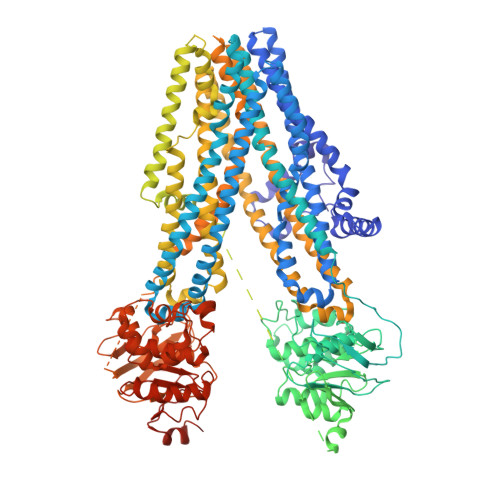
Molecular structures reveal synergistic rescue of Delta 508 CFTR by Trikafta modulators.
Fiedorczuk, K., Chen, J.(2022) Science 378: 284-290
- PubMed: 36264792 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.ade2216
- Primary Citation Related Structures:
8EIG, 8EIO, 8EIQ, 8EJ1 - PubMed Abstract:
The predominant mutation causing cystic fibrosis, a deletion of phenylalanine 508 (Δ508) in the cystic fibrosis transmembrane conductance regulator (CFTR), leads to severe defects in CFTR biogenesis and function. The advanced therapy Trikafta combines the folding corrector tezacaftor (VX-661), the channel potentiator ivacaftor (VX-770), and the dual-function modulator elexacaftor (VX-445). However, it is unclear how elexacaftor exerts its effects, in part because the structure of Δ508 CFTR is unknown. Here, we present cryo-electron microscopy structures of Δ508 CFTR in the absence and presence of CFTR modulators. When used alone, elexacaftor partially rectified interdomain assembly defects in Δ508 CFTR, but when combined with a type I corrector, did so fully. These data illustrate how the different modulators in Trikafta synergistically rescue Δ508 CFTR structure and function.
- Laboratory of Membrane Biology and Biophysics, The Rockefeller University, New York, NY 10065, USA.
Organizational Affiliation: