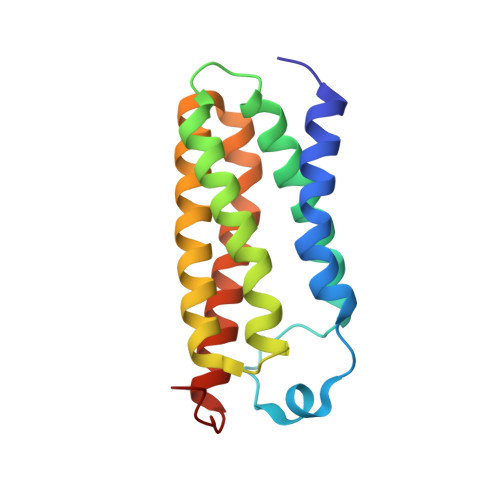
New TSPO Crystal Structures of Mutant and Heme-Bound Forms with Altered Flexibility, Ligand Binding, and Porphyrin Degradation Activity.
Liu, J., Hiser, C., Li, F., Hall, R., Garavito, R.M., Ferguson-Miller, S.(2023) Biochemistry 62: 1262-1273
- PubMed: 36947867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.2c00612
- Primary Citation Related Structures:
8E7W, 8E7X, 8E7Y, 8E7Z - PubMed Abstract:
The ancient protein TSPO (translocator protein 18kD) is found in all kingdoms and was originally identified as a binding site of benzodiazepine drugs. Its physiological function remains unclear, although porphyrins are conserved ligands. Several crystal structures of bacterial TSPO and nuclear magnetic resonance structures of a mouse form have revealed monomer and dimer configurations, but there have been no reports of structures with a physiological ligand. Here, we present the first X-ray structures of Rhodobacter sphaeroides TSPO with a physiological ligand bound. Two different variants (substituting threonine for alanine at position 139 (A139T) and phenylalanine for alanine at position 138 (A138F)) yielded well-diffracting crystals giving structures of both apo- and heme-containing forms. Both variants have wild-type micromolar affinity for heme and protoporphyrin IX, but A139T has very low ability to accelerate the breakdown of porphyrin in the presence of light and oxygen. The binding of heme to one protomer of the dimer of either mutant induces a more rigid structure, both in the heme-binding protomer and the protomer without heme bound, demonstrating an allosteric response. Ensemble refinement of the X-ray data reveals distinct regions of altered flexibility in response to single heme binding to the dimer. The A139T variant shows a more rigid structure overall, which may relate to extra hydrogen bonding of waters captured in the heme crevice. As TSPO has been suggested to have a role in heme delivery from mitochondria to the cytoplasm, the new structures provide potential clues regarding the structural basis of such activity.
- Department of Chemistry, Michigan State University, East Lansing, Michigan 48824, United States.
Organizational Affiliation: